Chapter 7 Deeper Understanding
In this chapter I consider some hypotheses that can be tested with relatively simple methods.
7.1 Regression Discontinuity
This subsection will show a fun test that we can examine in real time. Some governors of US states have decided to reopen businesses much sooner than recommended by public health officials. For example, Georgia has decided to reopen gyms, massage therapy, tatoo shops, barbers and hair stylists on April 24th with restaurants, theaters and private social clubs opening up on the 27th. It will be difficult to practice social distancing while getting a tatoo or a body piercing.
Pending: I’ll plot the data for Georgia by county 3 weeks before the 24th and 3 weeks after. It will take a few days, possibly up to 14 days, to see the effects on the positive testing rate due to a higher rate of social contact, and maybe up to two weeks to see any effects on the death rate. I’ll examine a similar plot for an adjacent state that has not relaxed their social distancing mandate for comparison.
This example will illustrate the use of the regression discontinuity (RD) design, though before the causality police get on my case for not clarifying my assumptions I’ll point out that I won’t be using this to claim I have estimated the causal effect of social distancing. I’ll illustrate the idea of this design, its characteristic plot and point to papers where the reader can learn about the various approaches to extracting additional information from this type of research design.
The city of Atlanta is in Fulton county, which is the top curve
datacov.US.long <- gather(datacov.US, day, count, `1/22/20`:(names(datacov.US)[ncol(datacov.US)]))
datacov.US.long$day <- as.Date(datacov.US.long$day, "%m/%d/%y")
datacov.US.long$day.numeric <- as.numeric(datacov.US.long$day) -
18283
# drop two counties because unassigned or out of GA;
# total 159 counties per wiki
# https://en.wikipedia.org/wiki/List_of_counties_in_Georgia
georgia.df <- subset(datacov.US.long, `Province/State` ==
"Georgia") %>% filter(!Admin2 %in% c("Out of GA", "Unassigned"))
# all county plot ggplot(subset(georgia.df, day >
# as.Date('2020/04/1')), aes(day,count, group=Admin2,
# color=Admin2)) + geom_line() + ggtitle('Georgia
# counties')
# focus just on counties with cases > 100 as of the
# compilation of the report red vertical line at 4/24
# and blue vertical line at 4/27
georgia.df.max <- georgia.df %>% group_by(Admin2) %>% mutate(maxcount = max(count,
na.rm = T)) %>% ungroup()
range.ylim <- range(georgia.df.max$maxcount)
# add two regression lines to plot before/after cutoff
georgia.df.max$cutoff <- factor(ifelse(georgia.df.max$day <=
as.Date("2020/04/24"), "before", "after"))
subset(georgia.df.max, day > as.Date("2020/04/1") & maxcount >=
1000) %>% mutate(label = if_else(day.numeric == max(day.numeric),
as.character(Admin2), NA_character_)) %>% ggplot(aes(day,
count, group = Admin2, color = Admin2)) + geom_line() +
ggtitle("Georgia counties with greater than 1000 cases") +
geom_vline(xintercept = as.Date("2020/04/24"), color = "red") +
ylim(range.ylim) + theme_bw() + geom_label_repel(aes(label = label),
nudge_x = 2, na.rm = T, segment.color = "grey90", label.size = 0.01,
size = 2.5, show.legend = F) + scale_x_date(limits = c(as.Date("2020/04/1"),
as.Date(max.day + 5))) + theme(legend.position = "none",
plot.margin = margin(0.1, 1, 0.1, 0.1, "cm")) #+
# stat_smooth(data=subset(georgia.df.max,
# Admin2=='Fulton'),aes(x=day,y=count,group=cutoff),
# method='lm')For comparison, I’ll pick an adjacent state that has not relaxed their restrictions. Alabama seems a safe comparison as both South Carolina and Florida have begin partial openings (e.g., Florida opened beaches prior to April 24). One could also examine Tennessee and North Carolina, which also border with Georgia.
# drop two counties because unassigned or out of GA;
# total 159 counties per wiki
# https://en.wikipedia.org/wiki/List_of_counties_in_Georgia
alabama.df <- subset(datacov.US.long, `Province/State` ==
"Alabama") %>% filter(!Admin2 %in% c("Out of AL", "Unassigned"))
# all county plot ggplot(subset(alabama.df, day >
# as.Date('2020/04/1')), aes(day,count, group=Admin2,
# color=Admin2)) + geom_line() + ggtitle('Alabama
# counties')
# focus just on counties with cases > 1000 as of the
# compilation of the report red vertical line at 4/24
# and blue vertical line at 4/27
alabama.df.max <- alabama.df %>% group_by(Admin2) %>% mutate(maxcount = max(count,
na.rm = T)) %>% ungroup()
subset(alabama.df.max, day > as.Date("2020/04/1") & maxcount >=
1000) %>% mutate(label = if_else(day.numeric == max(day.numeric),
as.character(Admin2), NA_character_)) %>% ggplot(aes(day,
count, group = Admin2, color = Admin2)) + geom_line() +
ggtitle("Alabama counties with greater than 1000 cases") +
geom_vline(xintercept = as.Date("2020/04/24"), color = "red") +
ylim(range.ylim) + theme_bw() + geom_label_repel(aes(label = label),
nudge_x = 2, na.rm = T, segment.color = "grey90", label.size = 0.01,
size = 2.5, show.legend = F) + scale_x_date(limits = c(as.Date("2020/04/1"),
as.Date(max.day + 5))) + theme(legend.position = "none",
plot.margin = margin(0.1, 1, 0.1, 0.1, "cm"))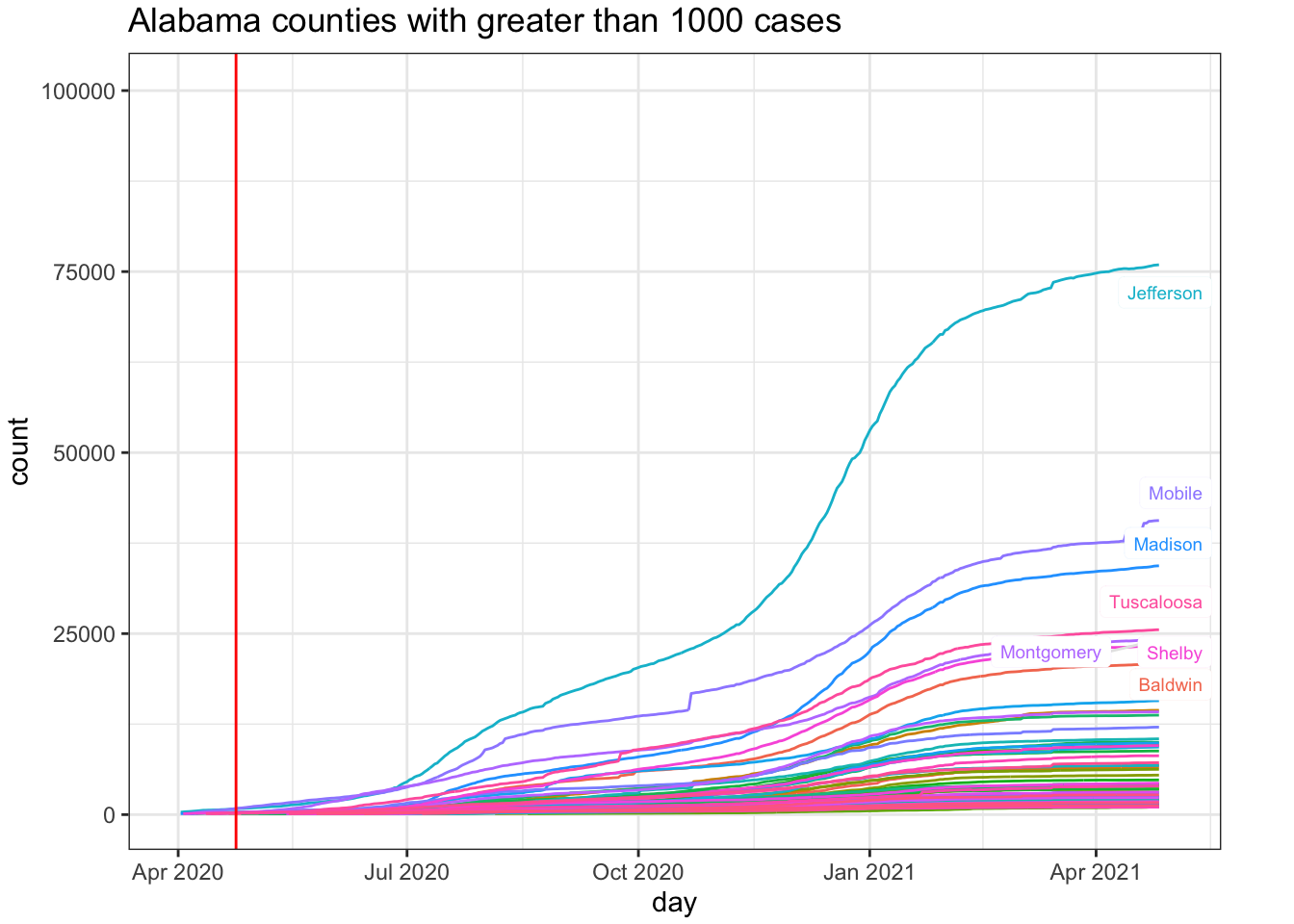
While I have a preference for examining incidence plots, the county-level data are too noisy to make incidence plots useful. But, at least, they do highlight the noise unlike the cumulative count data that acts like a visual smoother to the data patterns in terms of the visual effect on the curves. Here is the incidence plot for Fulton county in Georgia. The red vertical line marks April 24, 2020, when many restrictions in Georgia were lifted. For comparison, I also show Montgomery county in Alabama. We see that the trend remains the same for Fulton county George post restrictions but the comparison county in Alabama exhibits a higher rate of Covid-19 cases. This is opposite to hypothesis.
county.level <- georgia.df %>% subset(Admin2 == "Fulton" &
day > as.Date("2020/03/11")) %>% mutate(daily = c(NA,
diff(count)))
ggplot(county.level, aes(x = day, y = daily)) + geom_bar(stat = "identity") +
scale_x_date(date_breaks = "1 week", date_labels = "%d %b") +
labs(y = "Daily incremental incidence", title = "Fulton County, Georgia: Positive US Covid-19 Cases") +
theme(legend.position = "none", strip.text.y = element_text(size = 11),
axis.text.x = element_text(angle = 90, vjust = 0.5,
hjust = 1)) + geom_vline(xintercept = as.Date("2020/04/24"),
color = "red")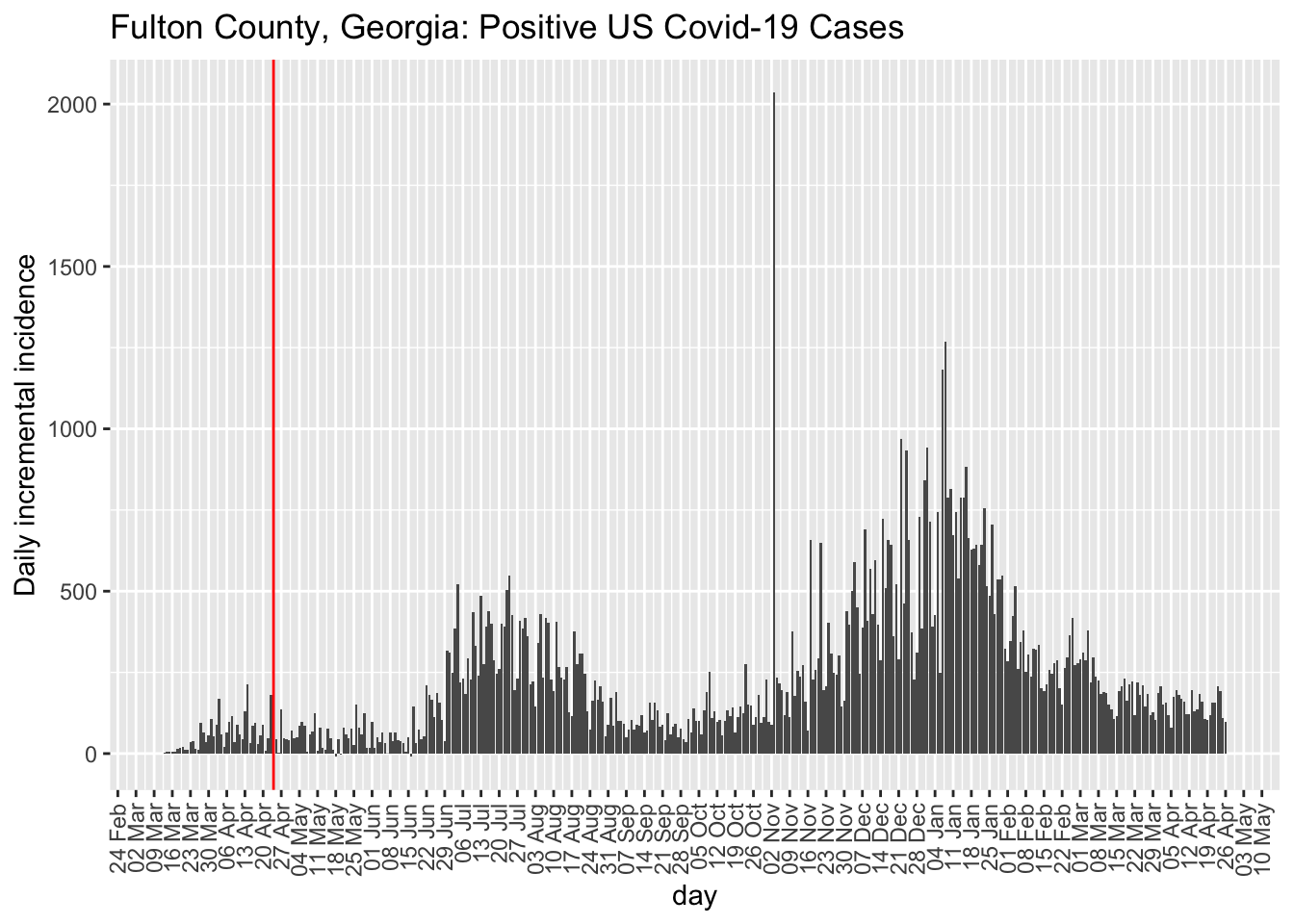
county.level <- alabama.df %>% subset(Admin2 == "Montgomery" &
day > as.Date("2020/03/11")) %>% mutate(daily = c(NA,
diff(count)))
ggplot(county.level, aes(x = day, y = daily)) + geom_bar(stat = "identity") +
scale_x_date(date_breaks = "1 week", date_labels = "%d %b") +
labs(y = "Daily incremental incidence", title = "Montgomery County, Alabama: Positive US Covid-19 Cases") +
theme(legend.position = "none", strip.text.y = element_text(size = 11),
axis.text.x = element_text(angle = 90, vjust = 0.5,
hjust = 1)) + geom_vline(xintercept = as.Date("2020/04/24"),
color = "red")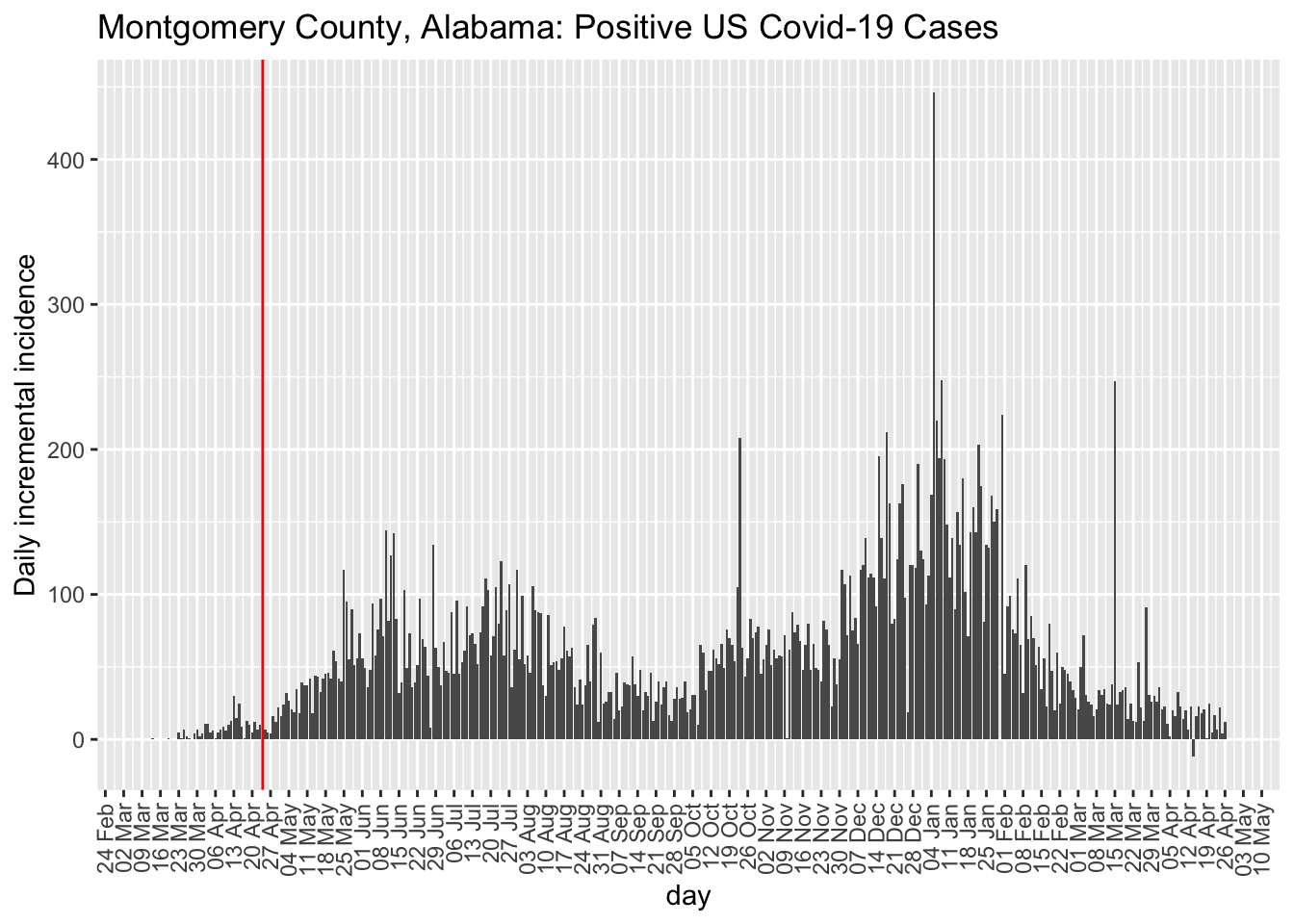
7.2 Hospital Readmission Data
My colleague Jack Iwashyna, who you will learn more about in the concluding chapter, suggested a hypothesis in mid April 2020. Research suggests that 40% of ARDS patients are re-admitted to the hospital within 1 year of discharge. This estimate came from a national, multicenter study with 839 ARDS survivors. Most covid-19 deaths are related to ARDS (more on this in the concluding chapter). Granted traditional ARDS patients may be different than covid-19-related ARDS patients but let’s use this as a plausible estimate of hospital re-admission. If we will see roughly a 40% chance of hospital readmission, it means that we may see additional hospitalizations in the coming months on top of the already steady stream of new covid-19 hospitalizations. And in the fall, if the predicted “second covid-19 wave” hits that could be compounded by additional hospital readmissions of people recently discharged or currently in the hospital. As the months progress, I’ll present analyses examining this hypothesis along with models that will assess second waves.
The good news is that the study found that patient-reported physical activity and quality of life status is associated with fewer re-admissions. So if we can get these covid-19-related ARDS survivors walking and exercising we may have some hope. Unfortunately, many ARDS survivors have trouble merely standing up from a chair let alone going for a walk around the block wearing a mask.
More on this to come. Here are some initial plots but the data are suspect with various missing days and implausible values such as New York reporting 9 consecutive days of reporting 5016, which suggests lack of daily update.
Format data
# use covid tracking data set; start at march 9 and
# create day.numeric as before
covid.tracking.new <- covid.tracking %>% subset(date > as.Date("2020-03-18",
"%Y-%m-%d"))
# reorder the data.frame
covid.tracking.new <- arrange(covid.tracking.new, date,
group_by = state)
covid.tracking.new$day.numeric <- as.numeric(covid.tracking.new$date) -
18330
# clean up death, hospitalization, ICU and vent columns
# temporary fix on 4/27/20 one missing code came up so
# I'll set it to 0
covid.tracking.new$deathIncrease <- ifelse(is.na(covid.tracking.new$deathIncrease),
0, covid.tracking.new$deathIncrease)
# create separate dataframe for US totals
day.death.totals.US <- covid.tracking.new %>% group_by(date) %>%
summarise(daytotal = sum(deathIncrease, na.rm = T),
.groups = "drop")
# add population info
temp1 <- str_sub(allstates.long.new$prettyval[1:50], start = -2,
end = -1)
temp2 <- allstates.long.new$population[1:50]
ord <- order(temp1)
temp1 <- temp1[ord]
temp2 <- temp2[ord]
n <- table(covid.tracking.new$state)[1]
covid.tracking.new$population <- rep(temp2, n)
# compute per capita for death
covid.tracking.new$deathIncrease.pc <- covid.tracking.new$deathIncrease/covid.tracking.new$population *
1e+06
covid.tracking.new$deathIncrease.pc[covid.tracking.new$deathIncrease.pc <
0] <- 0
# Drop one outlier in NJ. Y scale is free across states.
covid.tracking.new$deathIncrease.pc.nooutlier <- covid.tracking.new$deathIncrease.pc
covid.tracking.new$deathIncrease.pc.nooutlier[covid.tracking.new$deathIncrease.pc.nooutlier >
100] <- NA
day.hospitalizedCurrently <- covid.tracking.new %>% group_by(date) %>%
summarise(daytotal = sum(hospitalizedCurrently, na.rm = T),
.groups = "drop")
day.inIcuCurrently <- covid.tracking.new %>% group_by(date) %>%
summarise(daytotal = sum(inIcuCurrently, na.rm = T),
.groups = "drop")
day.onVentilatorCurrently <- covid.tracking.new %>% group_by(date) %>%
summarise(daytotal = sum(onVentilatorCurrently, na.rm = T),
.groups = "drop")ggplot(subset(covid.tracking, date > as.Date("2020/03/20")),
aes(date, hospitalizedCurrently, group = state, color = state)) +
geom_line() + ggtitle("Number Currently Hospitalized By State")
ggplot(subset(covid.tracking, date > as.Date("2020/03/20")),
aes(date, inIcuCurrently, group = state, color = state)) +
geom_line() + ggtitle("Number Currently in ICU By State")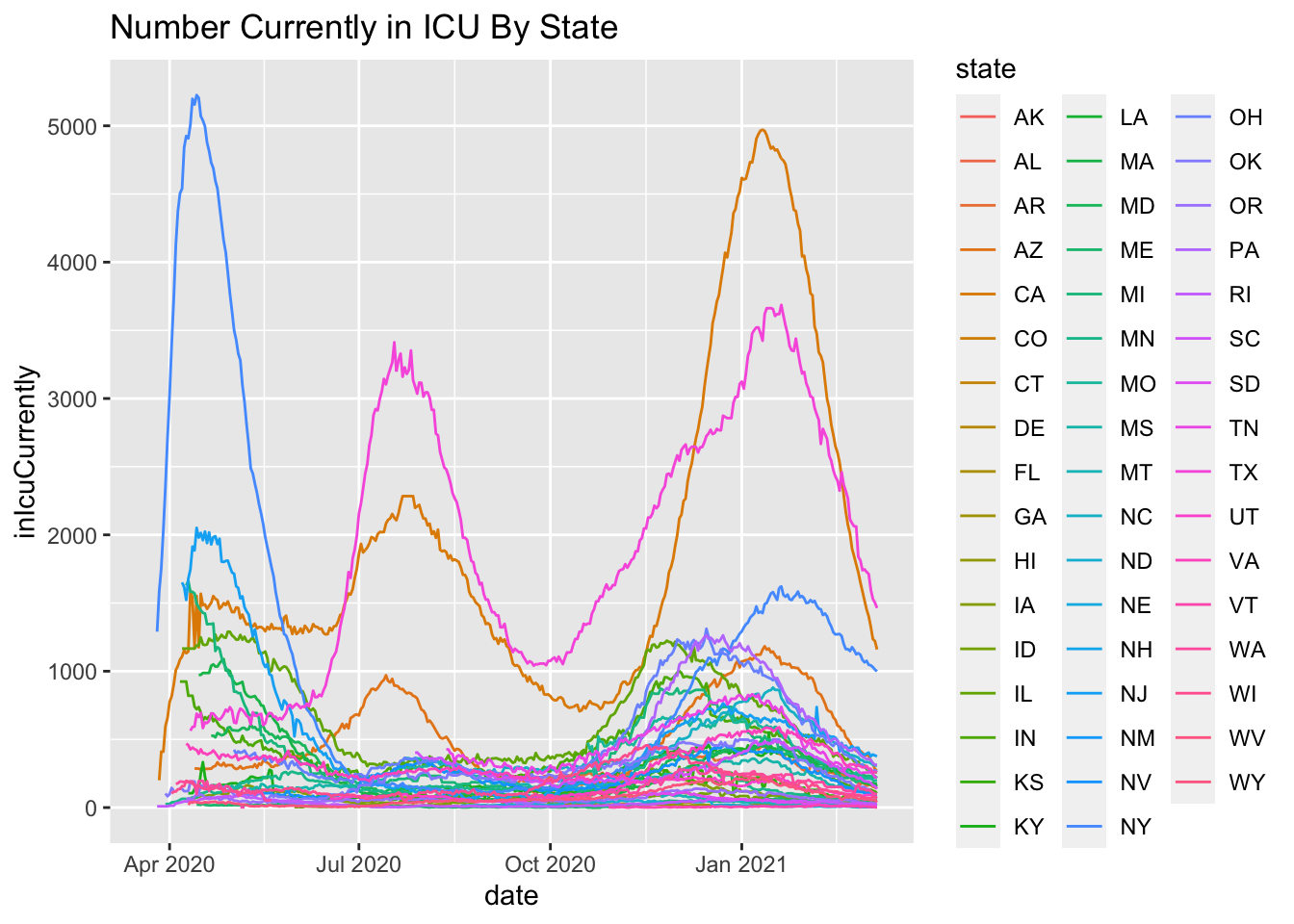
ggplot(subset(covid.tracking, date > as.Date("2020/03/20")),
aes(date, onVentilatorCurrently, group = state, color = state)) +
geom_line() + ggtitle("Number Currently on Ventilator By State")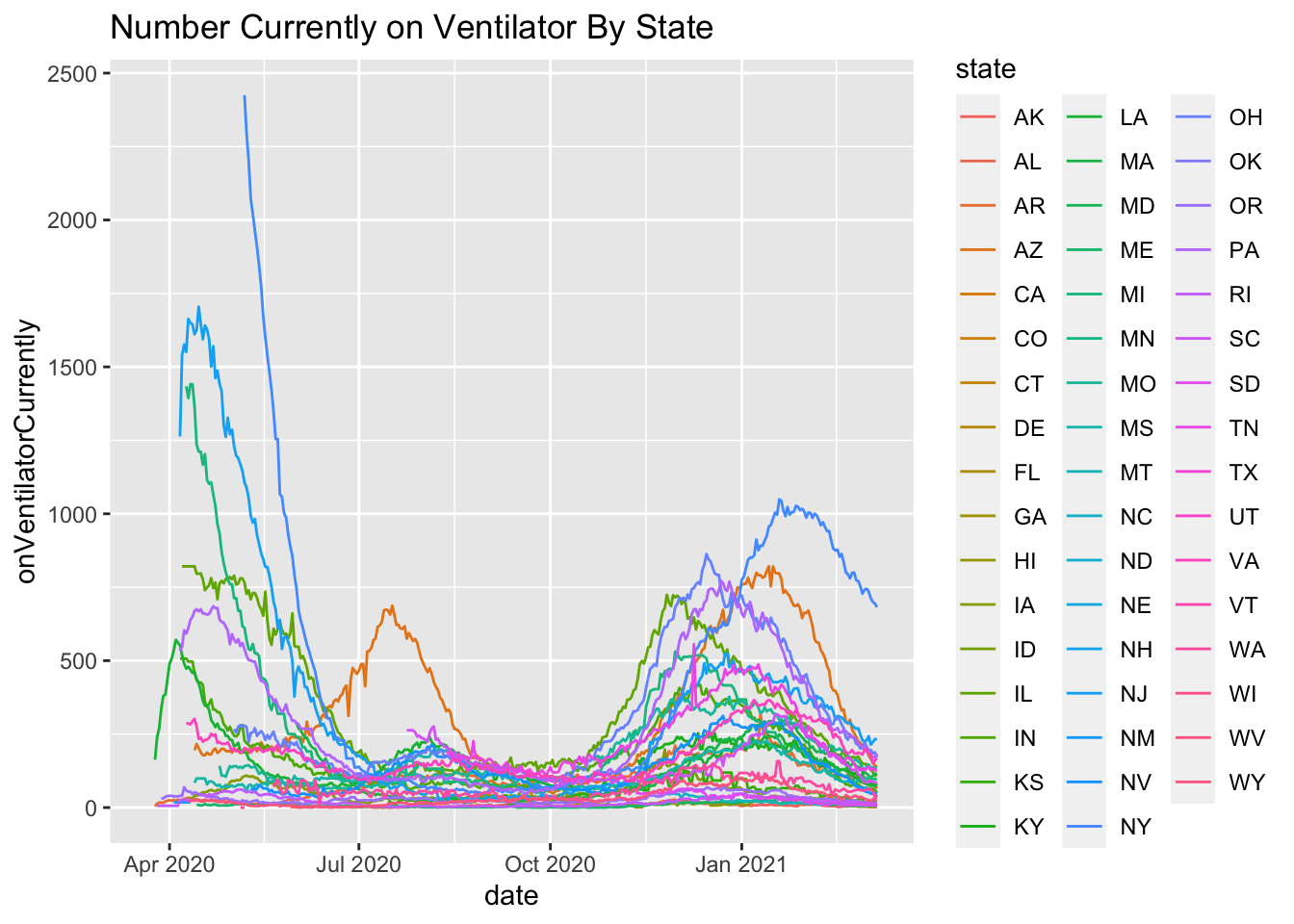
ggplot(day.hospitalizedCurrently, aes(date, daytotal)) +
geom_bar(stat = "identity", width = 1) + scale_x_date(date_breaks = "2 weeks",
date_labels = "%d %b", expand = expansion(mult = c(0.05,
0.15))) + labs(y = "Daily", title = "US Covid Hospitalization") +
theme(legend.position = "none", strip.text.y = element_text(size = 11),
axis.text.x = element_text(angle = 90, vjust = 0.5,
hjust = 1))
ggplot(day.inIcuCurrently, aes(date, daytotal)) + geom_bar(stat = "identity",
width = 1) + scale_x_date(date_breaks = "2 weeks", date_labels = "%d %b",
expand = expansion(mult = c(0.05, 0.15))) + labs(y = "Daily",
title = "US Covid ICU") + theme(legend.position = "none",
strip.text.y = element_text(size = 11), axis.text.x = element_text(angle = 90,
vjust = 0.5, hjust = 1))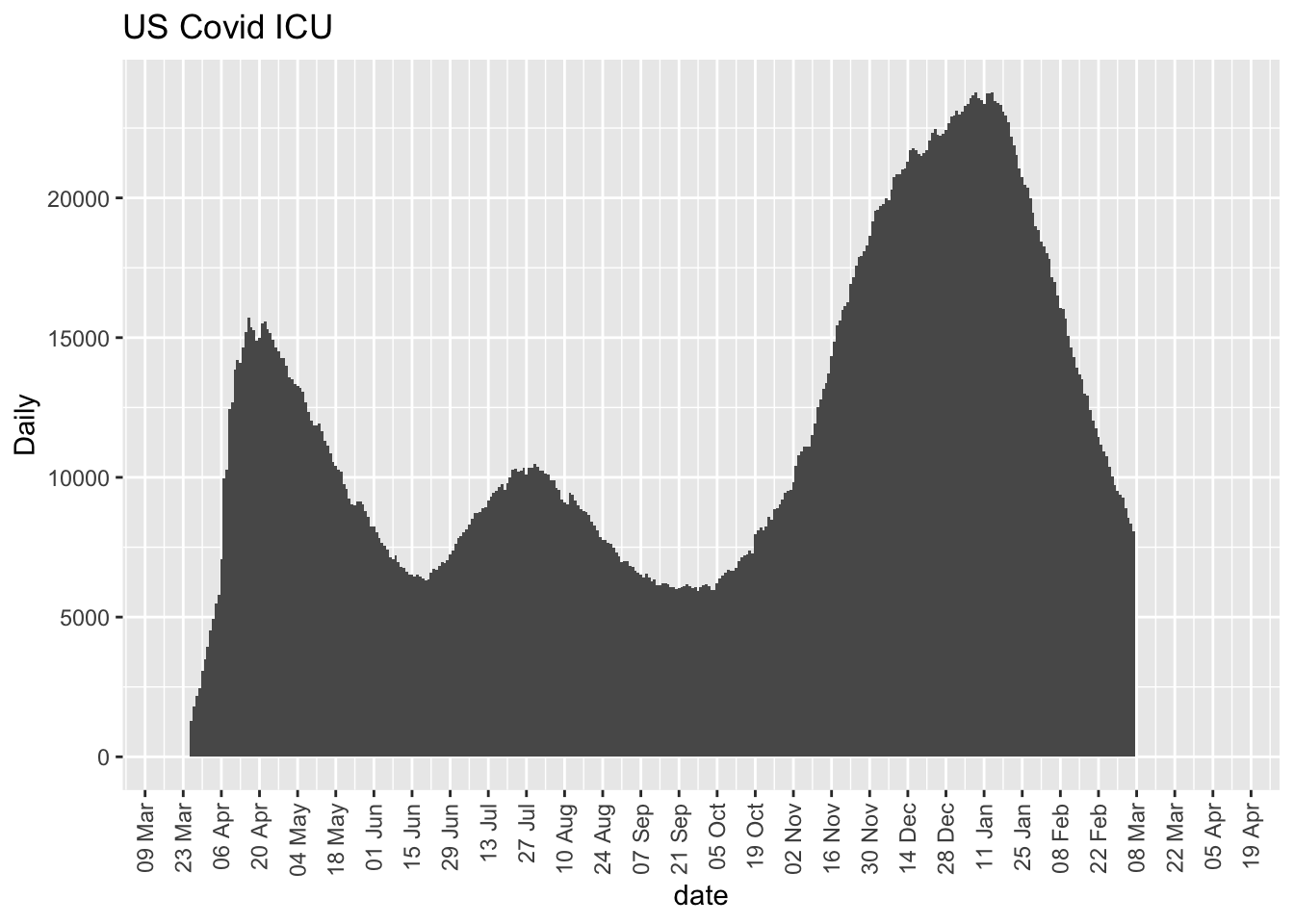
ggplot(day.onVentilatorCurrently, aes(date, daytotal)) +
geom_bar(stat = "identity", width = 1) + scale_x_date(date_breaks = "2 weeks",
date_labels = "%d %b", expand = expansion(mult = c(0.05,
0.15))) + labs(y = "Daily", title = "US Covid Ventilator") +
theme(legend.position = "none", strip.text.y = element_text(size = 11),
axis.text.x = element_text(angle = 90, vjust = 0.5,
hjust = 1))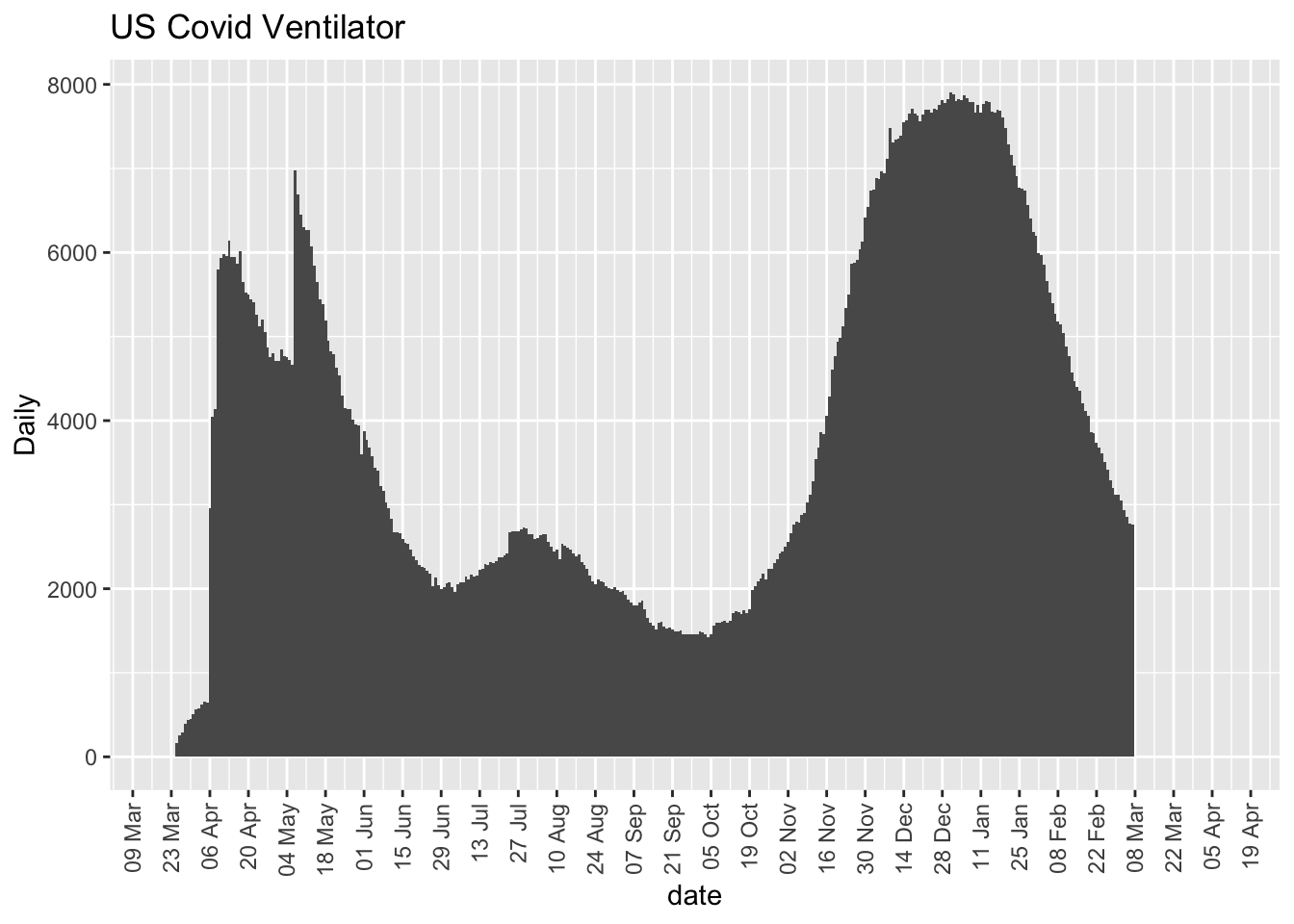
7.3 Deaths
The SIR model can be explored in more detail and its implications can be examined. Here I’ll focus on the simple SIR model without any of the bells-and-whistles I discussed in Chapter 6. The three primary equations of the SIR model are not easy to work with directly and they require numerical methods to approximate.
I’ll pursue one approach that avoids numerical methods by making some assumptions that will simplifying these expressions. As you will see, this simplification provides a reasonable fit to the data but their primary importance is in providing intuition into how to interpret these equations.
Following Keeling and Rohani (2008), if we assume that the \(R_0\) is relatively small and that people interact randomly, then the incidence curve from the SIR model can be approximated with this form
\[ a \;\;\mbox{sech}^2 (\kappa_0 + \kappa_1 t) \] where sech is the hyperbolic secant, its argument is a linear transformation of time t with a slope and intercept denoted by \(\kappa\)s, and parameter \(a\) is a multiplier. These parameters are each functions of the parameters of the SIR model (the \(\beta_1\) and \(\beta_2\) from Chapter 6) as well as the starting count at time 0. The hyperbolic secant can be reexpresssed in terms of exponentials: \(\mbox{sech} (x) = \frac{2}{\exp^{x} + \exp^{-x}}\). To arrive at this expression one could work with the SIR equations presented in Chapter 6 directly, but there is a simple way to get this form by assuming a Poisson distribution on the counts and showing that the probability that a randomly selected individual is not infected when the epidemic has an \(R_0\) is \(e^{\frac{-R_0}{N}}\) (i.e., a Poisson with k=0 and a rate of \(R_0/N\)); see Keeling and Rohani, 2008, Ch 2.
Let’s perform a direct fit to the daily death count, since death is a more appropriate outcome for the SIR model. For this section I’ll use the covid tracker data (see Chapter 3). We haven’t worked much with that data yet so I’ll also need to do some data cleaning and formatting to be consistent with the other analyses conducted in this book.
# define sir approximation to daily death count
richfu.sir <- function(d, a, b0, b1) {
a * sech(b1 * d + b0)^2
}Default smoothing parameters.
# incidence plot for US
ggplot(day.death.totals.US, aes(date, daytotal)) + geom_bar(stat = "identity",
width = 1) + scale_x_date(date_breaks = "2 weeks", date_labels = "%d %b",
expand = expansion(mult = c(0.05, 0.15))) + labs(y = "Daily",
title = "US Covid-19 Deaths") + theme(legend.position = "none",
strip.text.y = element_text(size = 11), axis.text.x = element_text(angle = 90,
vjust = 0.5, hjust = 1))
# incidence plot by state
ggplot(covid.tracking.new, aes(day.numeric, deathIncrease.pc.nooutlier)) +
geom_point(size = 1, color = "blue") + geom_smooth(color = "red",
size = 1) + facet_wrap_paginate(~state, nrow = 5, ncol = 5,
page = 1, scales = "free_y")
ggplot(covid.tracking.new, aes(day.numeric, deathIncrease.pc.nooutlier)) +
geom_point(size = 1, color = "blue") + geom_smooth(color = "red",
size = 1) + facet_wrap_paginate(~state, nrow = 5, ncol = 5,
page = 2, scales = "free_y")
# version was defined at the beginning of the the
# index.Rmd file get current tag and use a filename
# prefix
currentversion <- system("git describe --tags", intern = T)
save.image(file = paste0("data-archive/", currentversion,
"-dataworkspace.RData"))
print(paste0("File name associated with this run is ", currentversion,
"-dataworkspace.RData"))## [1] "File name associated with this run is v.95-123-g5a7aba6-dataworkspace.RData"# out <- nls(deathIncrease.pc ~ richfu.sir(day.numeric,
# a, b0,b1), data=covid.tracking.new,start=c(a=9,
# b0=2,b1=-0.1), algorithm='port',
# control=nls.control(maxiter=500))
# temporarily drop random b0 on 11/12 until convergence
# can be fixed
out <- nlme(deathIncrease.pc ~ richfu.sir(day.numeric, a,
b0, b1), fixed = a + b0 + b1 ~ 1, random = a + b1 ~
1, data = covid.tracking.new, start = c(a = 13, b0 = 1.75,
b1 = -0.01), verbose = T, groups = ~state, control = nlmeControl(msMaxIter = 400,
opt = "nlminb", pnlsTol = 0.01, msVerbose = T, minScale = 0.01,
maxIter = 400, tolerance = 0.5, msTol = 0.01))
# check public for recent run 8, .9, -.01, c(a=35,
# b0=1.23,b1=-.023)
# refit omitting the outlier out.nooutlier <-
# nlme(deathIncrease ~ richfu.sir(day.numeric, a,
# b0,b1), fixed = a+b0+b1~1, random = a+b0+b1~1,
# data=subset(covid.tracking.new,
# deathIncrease<800),start=c(a=40, b0=-1.8,b1=.03),
# verbose=T, groups=~state,
# control=nlmeControl(msMaxIter=1000,opt='nlminb',pnlsTol=.01,msVerbose=T,minScale=.01,
# maxIter=1000) )
# temporary kludge while I fix this, just use with
# outlier for nooutlier see chunk sir.approx.ny because
# have to drop subscript in temp.predict
out.nooutlier <- out
temp.data <- covid.tracking.new
# looks like two outliers as of 6/25/20, NY and NJ
# (prior only one NY)
dropped.data <- which(covid.tracking.new$deathIncrease >=
800)
starts <- nls(deathIncrease.pc ~ richfu.sir(day.numeric,
a, b0, b1), data = subset(covid.tracking.new, day.numeric <
43), start = c(a = 0.5, b0 = -2.7, b1 = 0.05), algorithm = "port")
out.first42days <- nlme(deathIncrease.pc ~ richfu.sir(day.numeric,
a, b0, b1), fixed = a + b0 + b1 ~ 1, random = a + b0 +
b1 ~ 1, data = subset(covid.tracking.new, day.numeric <
43), start = c(a = 13, b0 = -2.8, b1 = 0.074), verbose = T,
groups = ~state, control = nlmeControl(msMaxIter = 1000,
opt = "nlminb", pnlsTol = 0.01, msVerbose = T, minScale = 0.01,
maxIter = 1000, niterEM = 1000, msMaxEval = 1000,
tolerance = 1, msTol = 0.1))Play using Newton’s cooling equation; dropped one random effect due to convergence issues (come back to this later)
richfu.heat <- function(d, b0, b1, b2) {
b0 + b1 * exp(b2 * d)
}
# starts <- nls(deathIncrease.pc ~
# richfu.heat(day.numeric, b0,b1,b2),
# data=subset(covid.tracking.new,
# day.numeric>=34),start=c(b0=0.1, b1=30, b2=-.05),
# algorithm='port')
out.43days <- nlme(deathIncrease.pc ~ richfu.heat(day.numeric,
b0, b1, b2), fixed = b0 + b1 + b2 ~ 1, random = b0 +
b1 ~ 1, data = subset(covid.tracking.new, day.numeric >=
34), start = c(b0 = 7, b1 = 0.06, b2 = 0.02), verbose = T,
groups = ~state, control = nlmeControl(msMaxIter = 1000,
opt = "nlminb", pnlsTol = 0.001, msVerbose = T,
minScale = 0.001, maxIter = 1000, tolerance = 0.5,
msTol = 0.1))## 0: 129449.06: 0.602326 6.81877 -567.791
## 1: 129449.06: 0.602326 6.81877 -567.791
## 2: 129449.06: 0.602326 6.81877 -567.791
##
## **Iteration 1
## LME step: Loglik: -72745, nlminb iterations: 2
## reStruct parameters:
## state1 state2 state3
## 0.602 6.819 -567.791
## Beginning PNLS step: .. completed fit_nlme() step.
## PNLS step: RSS = 7669597
## fixed effects: 7.91 0.000944 0.0251
## iterations: 7
## Convergence crit. (must all become <= tolerance = 0.5):
## fixed reStruct
## 0.116 1.409
## 0: 129524.12: 1.06022 6.70106 1388.87
## 1: 129493.91: 0.0755384 6.87543 1388.87
## 2: 129490.20: 0.215146 6.94415 1388.87
## 3: 129488.19: 0.358290 7.00517 1388.87
## 4: 129488.04: 0.387607 7.01561 1388.87
## 5: 129487.98: 0.417966 7.04516 1388.83
## 6: 129487.97: 0.422310 7.02847 1388.83
## 7: 129487.96: 0.425856 7.02969 1388.77
## 8: 129487.92: 0.426733 7.03392 1387.87
## 9: 129487.24: 0.432724 7.05949 1373.54
## 10: 129481.01: 0.478578 7.24932 1179.17
## 11: 129474.89: 0.499143 7.29206 984.788
## 12: 129456.59: 0.501313 7.22853 211.345
## 13: 129452.87: 0.426045 7.09801 17.8588
## 14: 129451.94: 0.389207 7.00284 45.1323
## 15: 129451.86: 0.382587 6.96918 72.1348
## 16: 129451.86: 0.382785 6.96579 73.2912
## 17: 129451.86: 0.382769 6.96569 73.2091
##
## **Iteration 2
## LME step: Loglik: -72748, nlminb iterations: 17
## reStruct parameters:
## state1 state2 state3
## 0.383 6.966 73.209
## Beginning PNLS step: .. completed fit_nlme() step.
## PNLS step: RSS = 7578263
## fixed effects: 7.77 0.00538 0.0196
## iterations: 7
## Convergence crit. (must all become <= tolerance = 0.5):
## fixed reStruct
## 0.0184 299.3367
## 0: 129409.26: 0.425541 5.43352 -4.65539
## 1: 129407.39: 0.437279 5.15380 -4.65543
## 2: 129407.22: 0.434980 5.19003 -4.65544
## 3: 129407.17: 0.433832 5.21068 -4.65545
## 4: 129407.17: 0.433684 5.21626 -4.65545
## 5: 129407.16: 0.438147 5.22649 -4.65617
## 6: 129407.16: 0.433524 5.22633 -4.65619
## 7: 129407.16: 0.433524 5.22633 -4.65619
##
## **Iteration 3
## LME step: Loglik: -72703, nlminb iterations: 7
## reStruct parameters:
## state1 state2 state3
## 0.434 5.226 -4.656
## Beginning PNLS step: .. completed fit_nlme() step.
## PNLS step: RSS = 7507423
## fixed effects: 7.34 0.062 0.012
## iterations: 7
## Convergence crit. (must all become <= tolerance = 0.5):
## fixed reStruct
## 0.059 0.446## Nonlinear mixed-effects model fit by maximum likelihood
## Model: deathIncrease.pc ~ richfu.heat(day.numeric, b0, b1, b2)
## Data: subset(covid.tracking.new, day.numeric >= 34)
## AIC BIC logLik
## 145486 145540 -72736
##
## Random effects:
## Formula: list(b0 ~ 1, b1 ~ 1)
## Level: state
## Structure: General positive-definite, Log-Cholesky parametrization
## StdDev Corr
## b0 11.840 b0
## b1 0.458 0.135
## Residual 19.599
##
## Fixed effects: b0 + b1 + b2 ~ 1
## Value Std.Error DF t-value p-value
## b0 7.34 1.698 16448 4.32 0.000
## b1 0.06 0.081 16448 0.76 0.445
## b2 0.01 0.002 16448 5.50 0.000
## Correlation:
## b0 b1
## b1 0.034
## b2 0.107 -0.594
##
## Standardized Within-Group Residuals:
## Min Q1 Med Q3 Max
## -3.7055 -0.1256 -0.0178 0.0636 35.9176
##
## Number of Observations: 16500
## Number of Groups: 50We see that this function (red curve) does a reasonable job of fitting the daily death counts (black points) in New York. There is noise in these data that this form of the SIR model cannot easily pickup. I show an additional estimation using only the data up through day 42 (in green). This curve shows that the functional form fits reasonably well up through the peak but does not capture the asymmetry in terms of the slow decline in the frequency of new cases. This suggests there are additional processes that this simple implementation of the SIR model does not capture, and provides a clue for additional modeling changes one could implement.
## Nonlinear mixed-effects model fit by maximum likelihood
## Model: deathIncrease.pc ~ richfu.sir(day.numeric, a, b0, b1)
## Data: covid.tracking.new
## AIC BIC logLik
## 155064 155118 -77525
##
## Random effects:
## Formula: list(a ~ 1, b1 ~ 1)
## Level: state
## Structure: General positive-definite, Log-Cholesky parametrization
## StdDev Corr
## a 51.91897 a
## b1 0.00719 0.07
## Residual 19.08770
##
## Fixed effects: a + b0 + b1 ~ 1
## Value Std.Error DF t-value p-value
## a 23.14 7.51 17648 3.1 0.0021
## b0 -1.63 0.03 17648 -54.7 0.0000
## b1 0.00 0.00 17648 2.8 0.0046
## Correlation:
## a b0
## b0 -0.112
## b1 -0.002 -0.018
##
## Standardized Within-Group Residuals:
## Min Q1 Med Q3 Max
## -11.0146 -0.1113 -0.0180 0.0716 37.0873
##
## Number of Observations: 17700
## Number of Groups: 50out.nlme.deaths <- out
out.nlme.deaths.nooutlier <- out.nooutlier
# add predicted values to the data frame
covid.tracking.new$predicted <- predict(out, level = 1)
temp.predict <- predict(out.nooutlier, level = 1)
# if (length(dropped.data)==1)
# covid.tracking.new$predicted.nooutlier <-
# c(temp.predict[1:(dropped.data-1)], NA,
# #temp.predict[(dropped.data):length(temp.predict)])
covid.tracking.new.nooutlier <- covid.tracking.new[-dropped.data,
]
# drop [-dropped.data,] from temp.predict when fixing
# outlier above
# covid.tracking.new.nooutlier$predicted.nooutlier <-
# temp.predict[-dropped.data]
# just first 42 days
covid.tracking.new$predicted.42 <- predict(out.first42days,
newdata = data.frame(state = covid.tracking.new$state,
day.numeric = covid.tracking.new$day.numeric), level = 1)
# after 42 days covid.tracking.new$predicted.43plus <-
# predict(out.after43days,
# #newdata=data.frame(state=covid.tracking.new$state,
# day.numeric=covid.tracking.new$day.numeric), level=1)
p <- ggplot(subset(covid.tracking.new, state == "NY"), aes(x = day.numeric,
y = deathIncrease.pc)) + # scale_x_date(date_breaks='1 week', date_labels = '%d
# %b') +
geom_line(aes(day.numeric, predicted), color = "red") +
geom_point() + # geom_line(data=subset(covid.tracking.new.nooutlier,
# state=='NY'), aes(day.numeric, predicted.nooutlier),
# color='blue') +
geom_line(aes(day.numeric, predicted.42), color = "green") +
# geom_line(aes(day.numeric, predicted.43plus),
# color='black') +
labs(y = "Daily incidence ", title = "Daily Covid-19 Deaths in New York (red curve all data estimate; blue curve omitting single outlier") +
theme(legend.position = "none", strip.text.y = element_text(size = 11))
p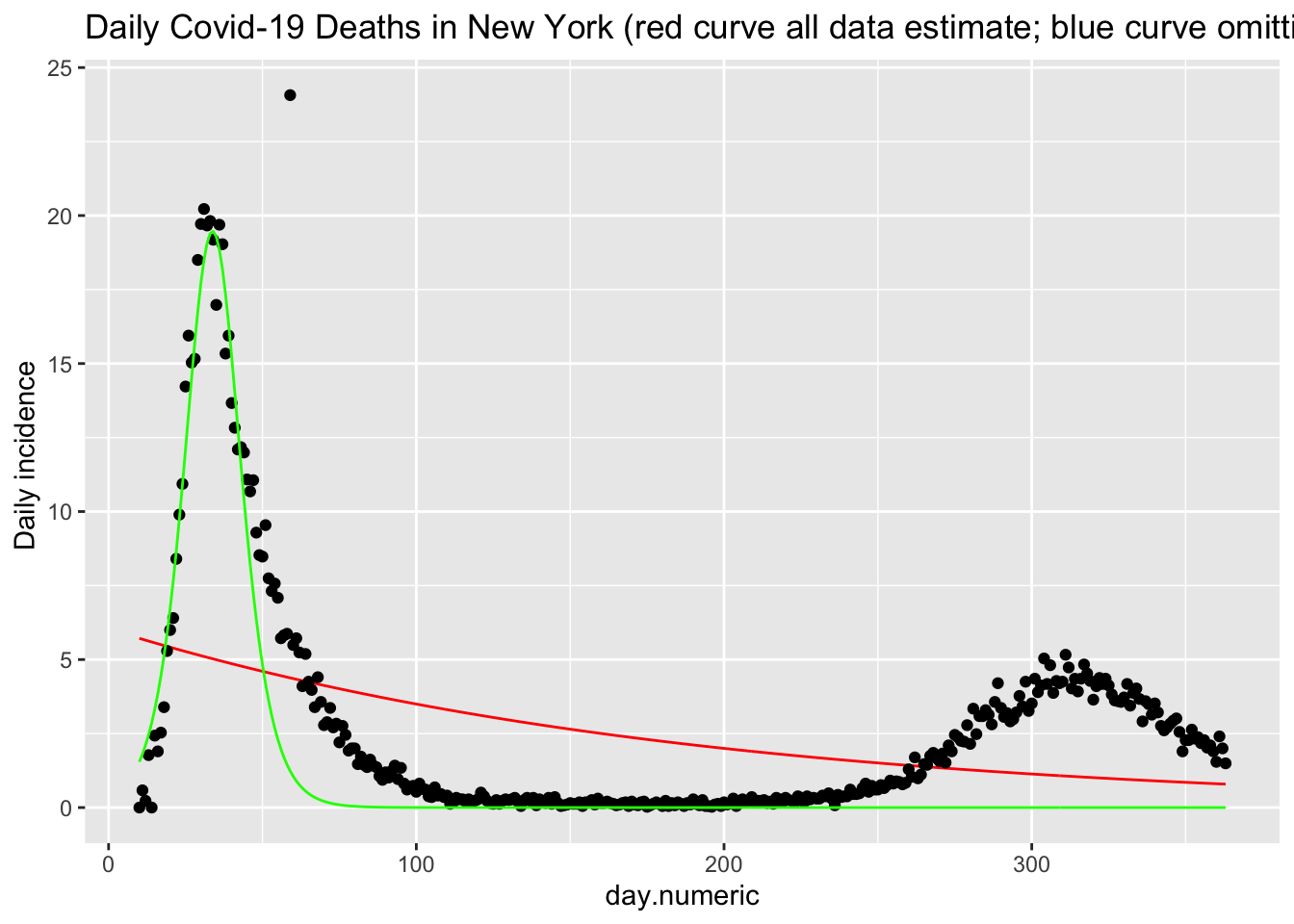
Add cooling
# 43days
covid.tracking.new$predicted.43 <- predict(out.43days, newdata = data.frame(state = covid.tracking.new$state,
day.numeric = covid.tracking.new$day.numeric), level = 1)
p + geom_line(data = subset(covid.tracking.new, state ==
"NY" & day.numeric >= 34), aes(day.numeric, predicted.43),
color = "black")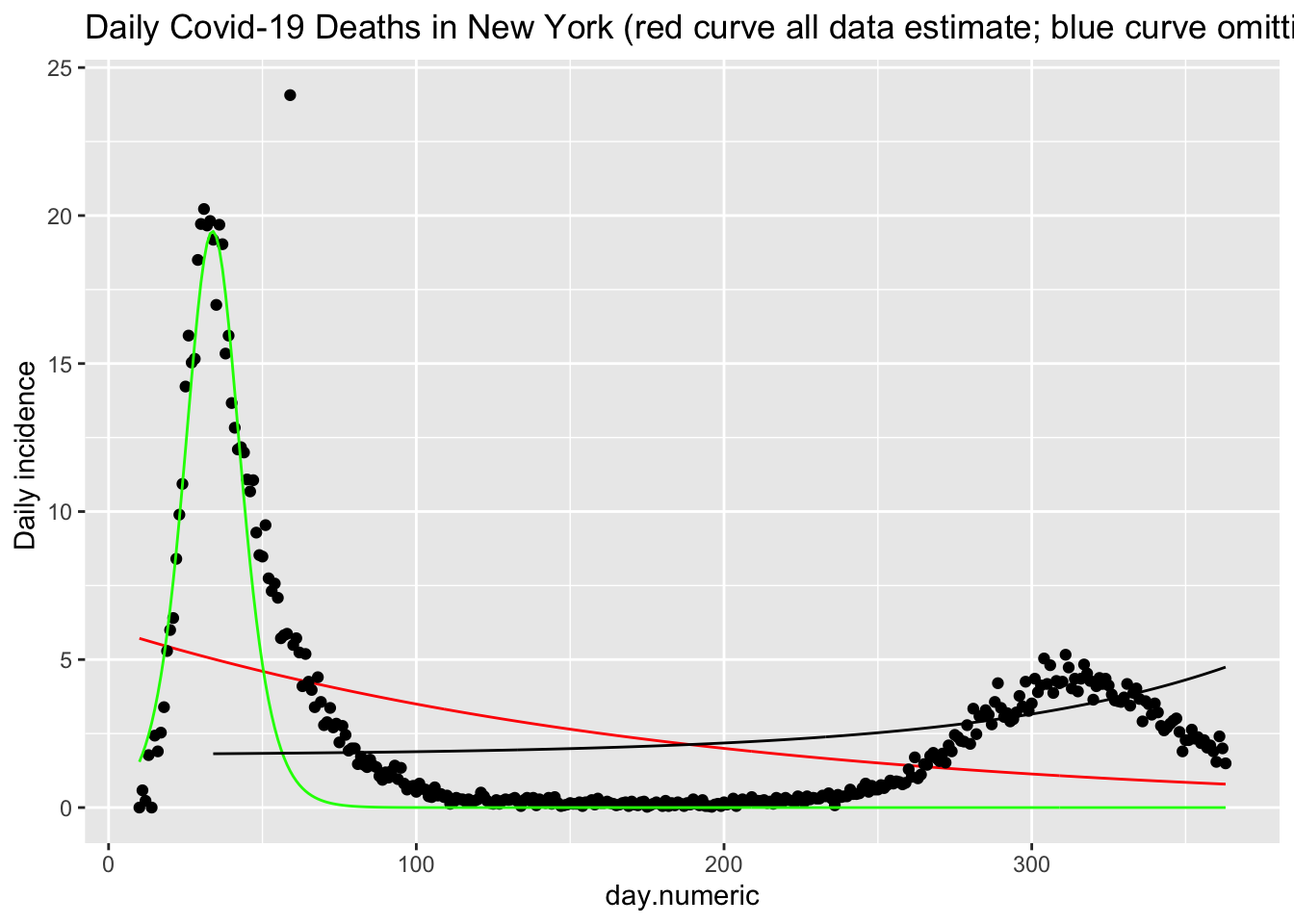
The data for death are quite noisy, for example, here are plots for both New Jersey and Michigan.
p1 <- ggplot(subset(covid.tracking.new, state == "NJ"),
aes(x = day.numeric, y = deathIncrease.pc)) + # scale_x_date(date_breaks='1 week', date_labels = '%d
# %b') +
geom_line(aes(day.numeric, predicted), color = "red") +
geom_point() + labs(y = "Daily incidence ", title = "Daily Covid-19 Deaths in New Jersey (per 1M)") +
theme(legend.position = "none", strip.text.y = element_text(size = 11))
p2 <- ggplot(subset(covid.tracking.new, state == "MI"),
aes(x = day.numeric, y = deathIncrease.pc)) + # scale_x_date(date_breaks='1 week', date_labels = '%d
# %b') +
geom_line(aes(day.numeric, predicted), color = "red") +
geom_point() + labs(y = "Daily incidence ", title = "Daily Covid-19 Deaths in Michigan (per 1M)") +
theme(legend.position = "none", strip.text.y = element_text(size = 11))
multiplot(p1, p2, cols = 1)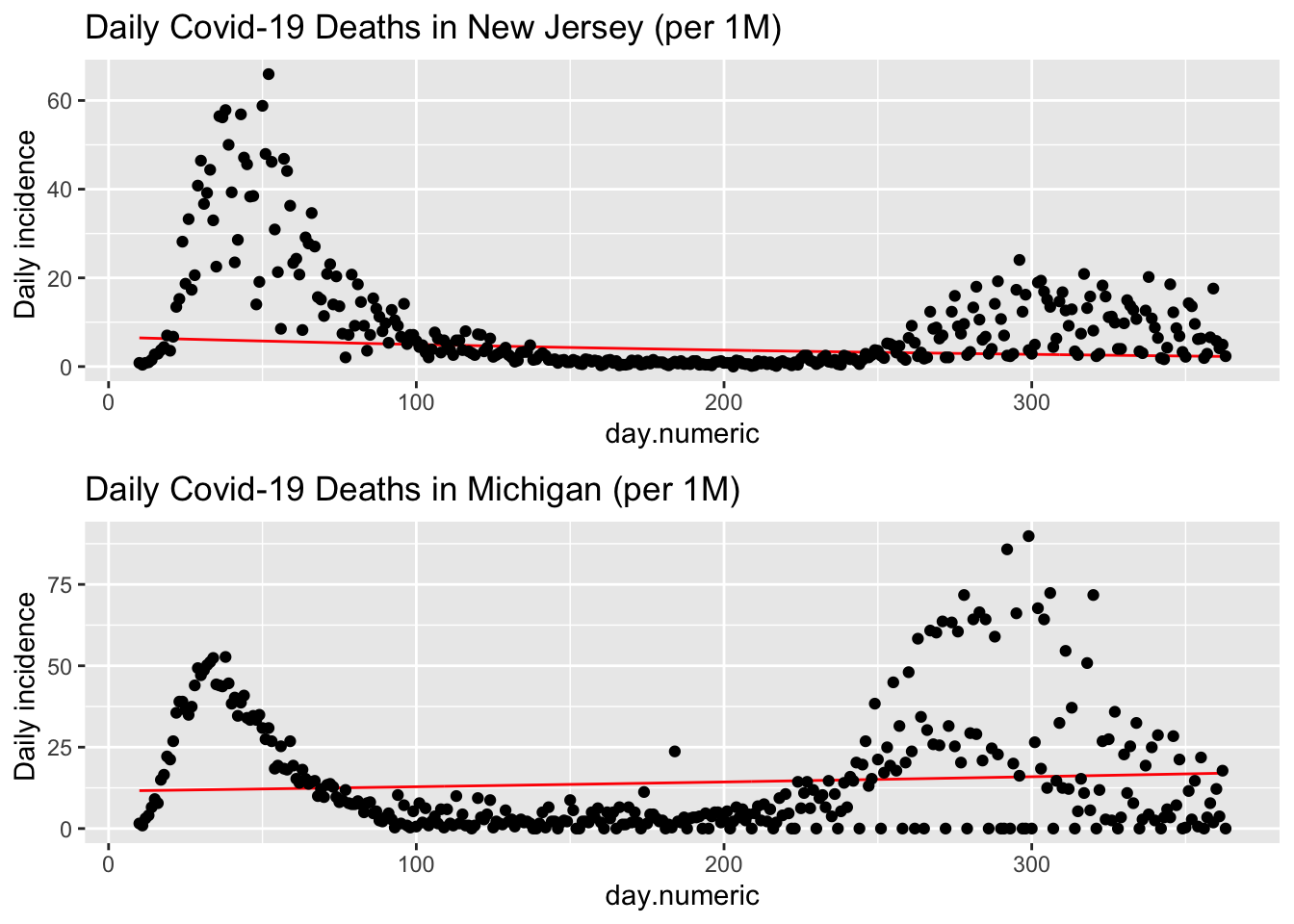
This large daily variability is troubling. One possibility is the way some states report deaths. In Michigan, if a death is determined to be covid-related it will be added to the day’s count even if the actual death occurred days earlier. To illustrate, on Saturday May 9, 2020, Michigan reported 133 deaths but 67 of those “deaths occurred in recent days” and determined to be covid-related after the fact were added to the Saturday total see. New York City, on the other hand, counts covid deaths when a positive test result and “probable” if the death certificate lists covid-19 as the cause of death (even if not verified with a known positive lab test). You may also be wondering about the single outlier for New York on May 6, 2020. That was due to how the state reconciled deaths from nursing home (see). Determining the number of deaths is not an easy task as this New York Times article shows. This also points out how we must examine data very carefully; our models and inferences will suffer greatly if we are too careless.
To be fair, these three plots (New York, New Jersey and Michigan) have different scales for the Y-axes, which may exaggerate the variability. I’ll redo the three plots with the same scale on the Y axis to check that possibility.
p1 <- ggplot(subset(covid.tracking.new, state == "NY"),
aes(x = day.numeric, y = deathIncrease.pc)) + # scale_x_date(date_breaks='1 week', date_labels = '%d
# %b') +
geom_line(aes(day.numeric, predicted), color = "red") +
geom_point() + labs(y = "Daily incidence ", title = "Daily Covid-19 Deaths in New York (per 1M)") +
theme(legend.position = "none", strip.text.y = element_text(size = 11)) +
ylim(0, 50)
p2 <- ggplot(subset(covid.tracking.new, state == "NJ"),
aes(x = day.numeric, y = deathIncrease.pc)) + # scale_x_date(date_breaks='1 week', date_labels = '%d
# %b') +
geom_line(aes(day.numeric, predicted), color = "red") +
geom_point() + labs(y = "Daily incidence ", title = "Daily Covid-19 Deaths in New Jersey (per 1M)") +
theme(legend.position = "none", strip.text.y = element_text(size = 11)) +
ylim(0, 50)
p3 <- ggplot(subset(covid.tracking.new, state == "MI"),
aes(x = day.numeric, y = deathIncrease.pc)) + # scale_x_date(date_breaks='1 week', date_labels = '%d
# %b') +
geom_line(aes(day.numeric, predicted), color = "red") +
geom_point() + labs(y = "Daily incidence ", title = "Daily Covid-19 Deaths in Michigan (per 1M)") +
theme(legend.position = "none", strip.text.y = element_text(size = 11)) +
ylim(0, 50)
multiplot(p1, p2, p3, cols = 1)
Here are all 50 states with the red curve indicating the best fitting curve. The simple model does not capture the second wave evident in states like LA and IA. Though does a solid job of capturing the more standard curves like MI, NY, and MD, and can pick up states with late single spikes such as TX and to some extent SC. The model is not picking up clear increases after day 150 in states like IL and MD.
ggplot(covid.tracking.new, aes(day.numeric, deathIncrease.pc.nooutlier)) +
geom_point(size = 1, color = "blue") + geom_line(aes(day.numeric,
predicted), color = "red") + facet_wrap_paginate(~state,
nrow = 5, ncol = 5, page = 1, scales = "free_y")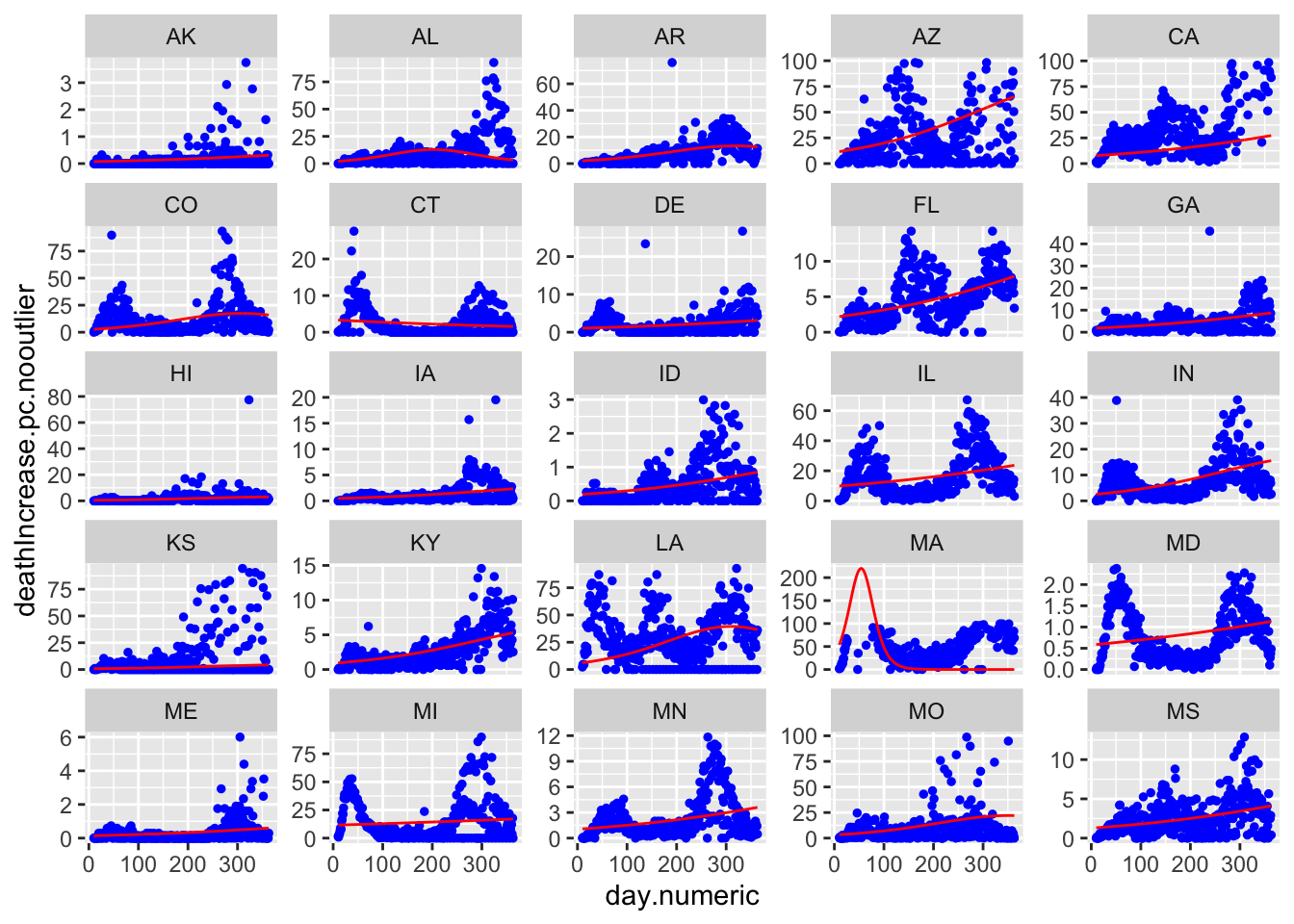
ggplot(covid.tracking.new, aes(day.numeric, deathIncrease.pc.nooutlier)) +
geom_point(size = 1, color = "blue") + geom_line(aes(day.numeric,
predicted), color = "red") + facet_wrap_paginate(~state,
nrow = 5, ncol = 5, page = 2, scales = "free_y")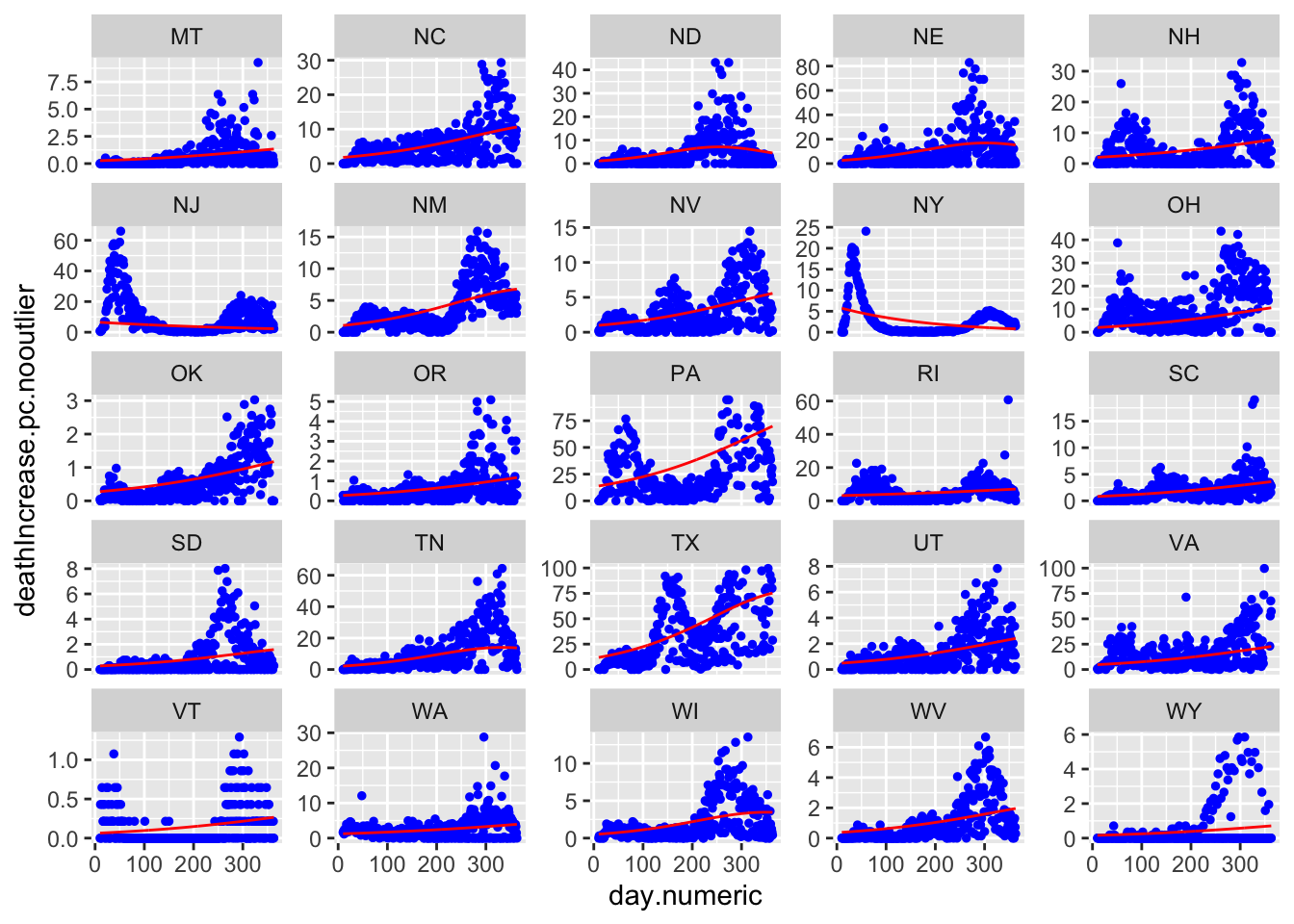
Predicting death counts.
nday = 10
predictdeath <- data.frame(day.numeric = rep(max.day.numeric:(max.day.numeric +
nday), 50), state = rep(rownames(coef(out.nlme.deaths.nooutlier)),
each = nday + 1))
predictdeath$prediction <- predict(out.nlme.deaths.nooutlier,
newdata = data.frame(day.numeric = predictdeath$day.numeric,
state = predictdeath$state), level = 1)
predictdeath$count <- predictdeath$prediction * rep(covid.tracking.new$population[1:50],
each = nday + 1)/1e+06
temp <- by(predictdeath$count, predictdeath$day.numeric,
sum)
temp## predictdeath$day.numeric: 413
## [1] 1889
## -----------------------------------------
## predictdeath$day.numeric: 414
## [1] 1891
## -----------------------------------------
## predictdeath$day.numeric: 415
## [1] 1893
## -----------------------------------------
## predictdeath$day.numeric: 416
## [1] 1895
## -----------------------------------------
## predictdeath$day.numeric: 417
## [1] 1897
## -----------------------------------------
## predictdeath$day.numeric: 418
## [1] 1899
## -----------------------------------------
## predictdeath$day.numeric: 419
## [1] 1901
## -----------------------------------------
## predictdeath$day.numeric: 420
## [1] 1903
## -----------------------------------------
## predictdeath$day.numeric: 421
## [1] 1905
## -----------------------------------------
## predictdeath$day.numeric: 422
## [1] 1907
## -----------------------------------------
## predictdeath$day.numeric: 423
## [1] 1908## [1] 20889## [1] 0FIX TO PERCAPITA DEATH
But a more interesting result emerges by examining this approximate functional form. We can integrate this approximation over t to see what form emerges for the cumulative death count. In this context, integration is analogous to summing the daily counts to create a cumulative count but we do this symbolically on the function rather than the data in order to learn the form of the function on the cumulative count.
The general form is
\[ \bigg[\frac{a}{\kappa_1 (1+e^{2(\kappa_0 + \kappa_1 t)} )}\bigg] \;\;\; \bigg[(e^{2(\kappa_0 + \kappa_1 t)} - 1)\bigg] + C \] The left square bracket term has the form of the logistic growth function and the right square bracket term has the form of an exponential growth function. The term \(C\) is the integration constant that we will set so that the cumulative sum starts at 0. Thus, this simplification of the SIR model yields a product of the two functions we explored in Chapter 5. It isn’t one or the other as we explored earlier, but this model suggests the curves approximately follow a combination of both forms. This is an example of the advantage of doing some mathematical modeling to be able to examine the implications of your model. In the earlier chapter we considered exponential and logistic growth models without careful attention to why we chose those functions. This type of mathematical modeling allows you to justify the functional forms you want to test based on implications from your hypotheses. Here I’m referring to the change equations of the SIR model as a set of hypotheses that govern how change occurs among the susceptibles, the infecteds and recovered. Those equations (with some additional assumptions and approximations) imply this product of both forms, which we can test and examine with data. Not counting the integration constant this approximation also has 3 parameters as we had in both the exponential form and the logistic growth form in Chapter 5.
# pull out the coefficients from the nlme
coef.ny <- coef(out)[rownames(coef(out)) == "NY", ]
a <- unlist(coef.ny[1])
k0 <- unlist(coef.ny[2])
k1 <- unlist(coef.ny[3])
t <- (covid.tracking.new$day.numeric[covid.tracking.new$state ==
"NY"])
pred <- a * (exp(2 * k1 * t + 2 * k0) - 1)/(k1 * (exp(2 *
k1 * t + 2 * k0) + 1))
# set integration constant so incidence at time 6 is 3
# (actual NY) pred <- pred + (-1*pred[length(pred)-5]) +
# 3 - 500 different approach for integration constant;
# need to check, seems ad hoc
const <- coef(lm(covid.tracking.new$death[covid.tracking.new$state ==
"NY"] - pred ~ 1))
pred <- pred + const
## compare to computed cumsum directly deathny <-
## rev(covid.tracking.new$death[covid.tracking.new$state=='NY'])
## predcumsum <-
## cumsum(rev(predict(out)[names(predict(out))=='NY']))
## cbind(deathny, pred,predcumsum)
preddat <- data.frame(day.numeric = t, day = covid.tracking.new$day.numeric[covid.tracking.new$state ==
"NY"], pred = pred)
ggplot(subset(covid.tracking.new, state == "NY"), aes(x = day.numeric,
y = death)) + geom_line(data = preddat, aes(day.numeric,
pred), color = "red") + geom_point() + labs(y = "Cumulative counts ",
title = "US Covid-19 Deaths in New York") + theme(legend.position = "none",
strip.text.y = element_text(size = 11))
The fit is ok given the simplifying assumptions that were made to derive this form, such as assuming that \(R_0\) is small, independence in transmission and some of the approximations that were used to derive this form (see Keeling & Rohani, 2008). We can reject a Poisson outright because the \(R_0\) value likely changed over time with different levels of social distancing.
It is helpful to estimate the logistic function directly on the cumulative death count and superimpose that curve (blue) on the same graph. We see that the pure logistic provides a better fit to these data. This shouldn’t be too surprising since I fit the logistic form directly to the data, whereas the red curve corresponding to the approximate SIR model takes into account other processes, has additional simplifying assumptions and approximations. Nonetheless, we can use the data and the fit to the models, to decide if we want to reject the SIR model because that model implies a functional form that fits worse than a simpler logistic growth model, use this finding to decide whether we should relax those simplifying assumptions and derive a new form, or use this finding to justify using the SIR model because the fit to the data is “close enough.” Of course, close enough depends on different applications. If one wants to describe a bunch of points with a simple curve or one wants to use the model to make forecasts. In the case of forecasting to inform public policy, then “close enough” better be pretty darn close because many people’s lives will be affected. Often, the disagreements between modelers about their predictions boils down to details of the assumptions each modeler makes and the kinds of simplifications they use.
7/10/20 dropped xmid random effect to get code to run; debug later, likely something weird with today’s data pull
# logistic on death; use same function as used for
# logistic growth on positive count data
covid.tracking.new$death.pc <- covid.tracking.new$death/covid.tracking.new$population
starts <- getInitial(death.pc ~ SSlogis(day.numeric, Asym = 1,
xmid = 1, scal = 1), data = covid.tracking.new)
tempdata <- covid.tracking.new[, c("day.numeric", "death.pc",
"state", "deathIncrease")]
out.logistic.death <- nlme(death.pc ~ SSlogis(day.numeric,
asym, xmid, scal), fixed = asym + xmid + scal ~ 1, random = xmid +
scal ~ 1, data = na.omit(tempdata), start = c(asym = starts[1],
xmid = starts[2], scal = starts[3]), verbose = T, groups = ~state,
control = nlmeControl(msMaxIter = 1000, opt = "nlminb",
pnlsTol = 0.001, msVerbose = T, minScale = 1, maxIter = 1000,
eval.mas = 1000, tolerance = 1))# add predicted values from the logistic regression;
# need to fix scaling
preddat$pred.logistic <- c((predict(out.logistic.death)[names(predict(out.logistic.death)) ==
"NY"]))
ggplot(subset(covid.tracking.new, state == "NY"), aes(x = day.numeric,
y = death)) + geom_line(data = preddat, aes(day.numeric,
pred), color = "red") + geom_point() + geom_line(data = preddat,
aes(day.numeric, pred.logistic), color = "blue")
labs(y = "Cumulative counts ", title = "US Covid-19 Deaths New York") +
theme(legend.position = "none", strip.text.y = element_text(size = 11))Compare AIC
(suppressed due to convergence issue)
Add phase plot of this model
# copy from logistic in previous chapter; edit to fit
# this function
logistic.pred.ts <- ts(diff(nydf.logistic$prediction))
plot(logistic.pred.ts, stats::lag(logistic.pred.ts, 1),
cex = 0.8, col = "blue", main = "Phase plot of New York Positive Counts for the Logistic Model",
xlab = "count(t)", ylab = "count(t+1)")
abline(0, 1)
lines(c(logistic.pred.ts), c(stats::lag(logistic.pred.ts,
1)), col = "red", type = "S")7.4 Functional Equations
Need to develop this section. Currently just a few plots. exponential is percentage but data and logistic are percap; need to make consistent.
# should pull out the coef from the fitted objects add
# the sech form
d <- 0:90
pred <- richfu(d, 0, 1.0288, 0.0578, 1)
test.1 <- data.frame(d = d, pred = c(NA, pred[-length(d)]),
pred.plus = c(NA, pred[-1]), group = rep(1, length(d)))
test.2 <- data.frame(d = d, pred = c(rep(NA, 2), pred[-seq(length(d) -
1, length(d), 1)]), pred.plus = c(rep(NA, 2), pred[-seq(1,
2, 1)]), group = rep(2, length(d)))
test.3 <- data.frame(d = d, pred = c(rep(NA, 3), pred[-seq(length(d) -
2, length(d), 1)]), pred.plus = c(rep(NA, 3), pred[-seq(1,
3, 1)]), group = rep(3, length(d)))
test.4 <- data.frame(d = d, pred = c(rep(NA, 4), pred[-seq(length(d) -
3, length(d), 1)]), pred.plus = c(rep(NA, 4), pred[-seq(1,
4, 1)]), group = rep(4, length(d)))
test.5 <- data.frame(d = d, pred = c(rep(NA, 5), pred[-seq(length(d) -
4, length(d), 1)]), pred.plus = c(rep(NA, 5), pred[-seq(1,
5, 1)]), group = rep(5, length(d)))
test <- rbind(test.1, test.2, test.3, test.4, test.5)
test$group <- factor(test$group)
ggplot(test, aes(x = pred, y = pred.plus, group = group,
color = group)) + geom_line() + xlab("f(t)") + ylab("f(t+k)") +
labs(color = "value of k")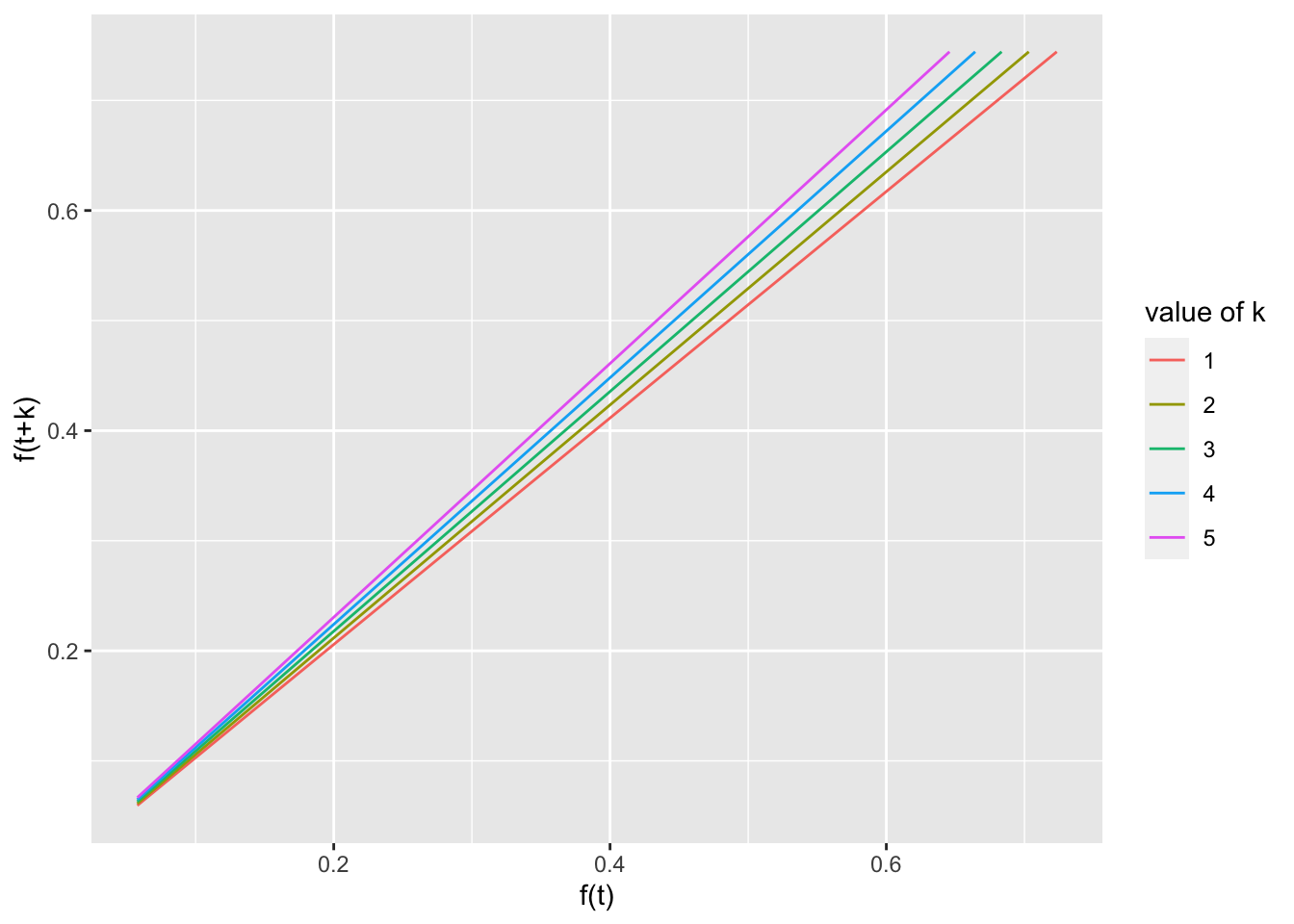
ggplot(data.frame(d, pred), aes(d, pred)) + geom_line() +
xlab("time") + ylab("count") + ggtitle("Best fitting exponential")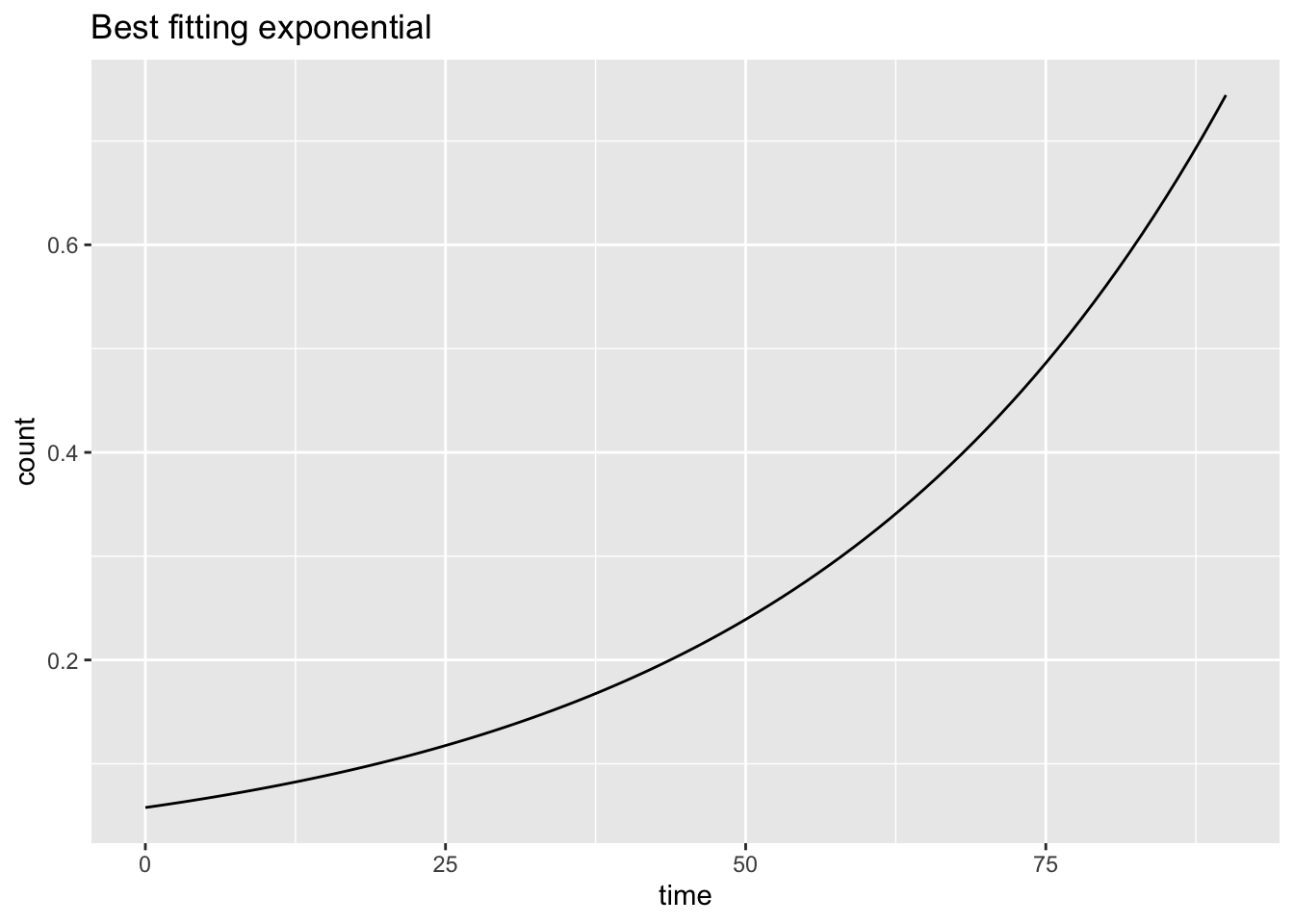
# redo for NY
d <- nydf.logistic$day.numeric
pred <- nydf.logistic$percap10000
test.1 <- data.frame(d = d, pred = c(NA, pred[-length(d)]),
pred.plus = c(NA, pred[-1]), group = rep(1, length(d)))
test.2 <- data.frame(d = d, pred = c(rep(NA, 2), pred[-seq(length(d) -
1, length(d), 1)]), pred.plus = c(rep(NA, 2), pred[-seq(1,
2, 1)]), group = rep(2, length(d)))
test.3 <- data.frame(d = d, pred = c(rep(NA, 3), pred[-seq(length(d) -
2, length(d), 1)]), pred.plus = c(rep(NA, 3), pred[-seq(1,
3, 1)]), group = rep(3, length(d)))
test.4 <- data.frame(d = d, pred = c(rep(NA, 4), pred[-seq(length(d) -
3, length(d), 1)]), pred.plus = c(rep(NA, 4), pred[-seq(1,
4, 1)]), group = rep(4, length(d)))
test.5 <- data.frame(d = d, pred = c(rep(NA, 5), pred[-seq(length(d) -
4, length(d), 1)]), pred.plus = c(rep(NA, 5), pred[-seq(1,
5, 1)]), group = rep(5, length(d)))
test <- rbind(test.1, test.2, test.3, test.4, test.5)
test$group <- factor(test$group)
ggplot(test, aes(x = pred, y = pred.plus, group = group,
color = group)) + geom_line() + xlab("f(t)") + ylab("f(t+k)") +
labs(color = "value of k")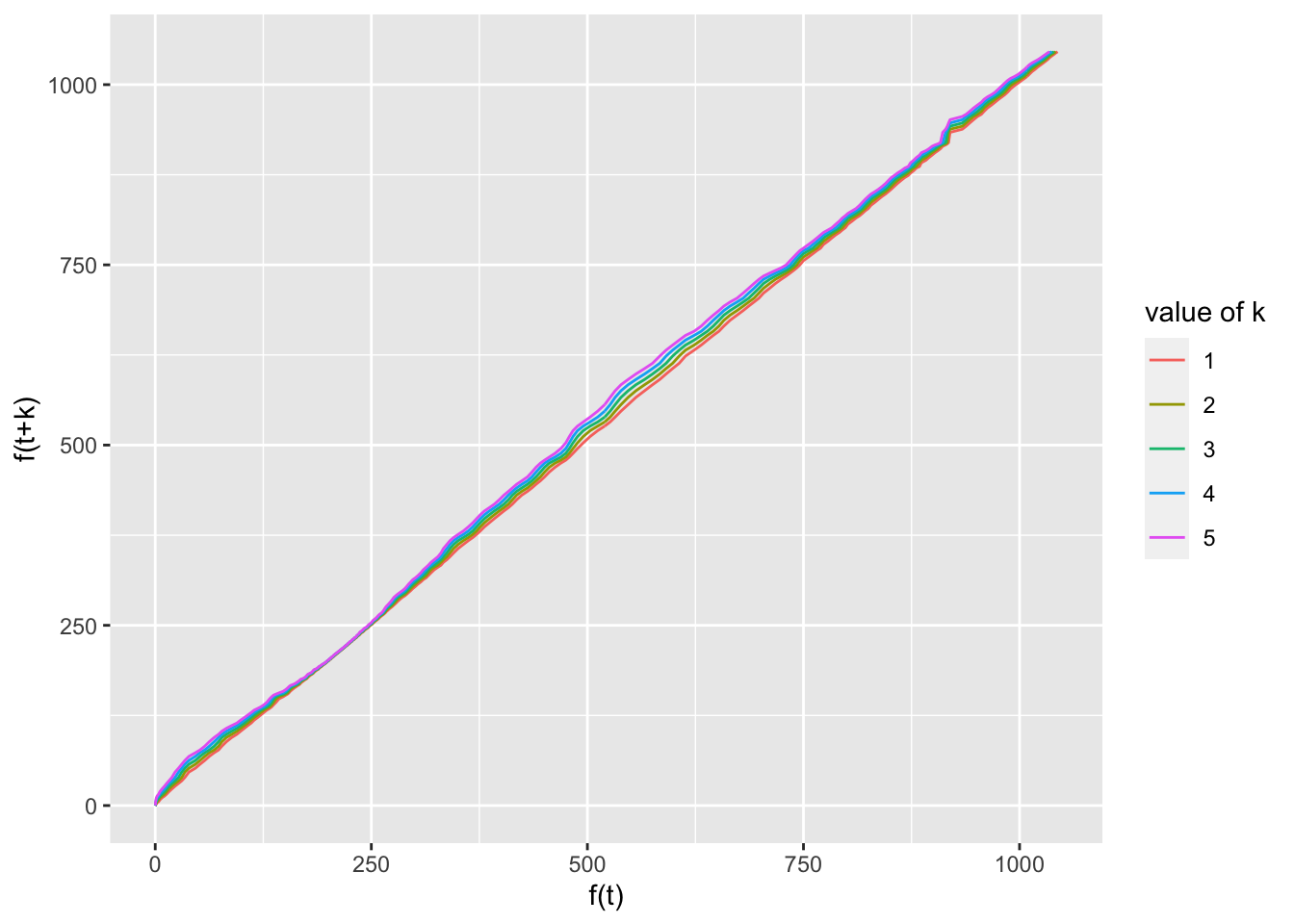
ggplot(data.frame(d, pred), aes(d, pred)) + geom_line() +
xlab("time") + ylab("count") + ggtitle("Actual Data")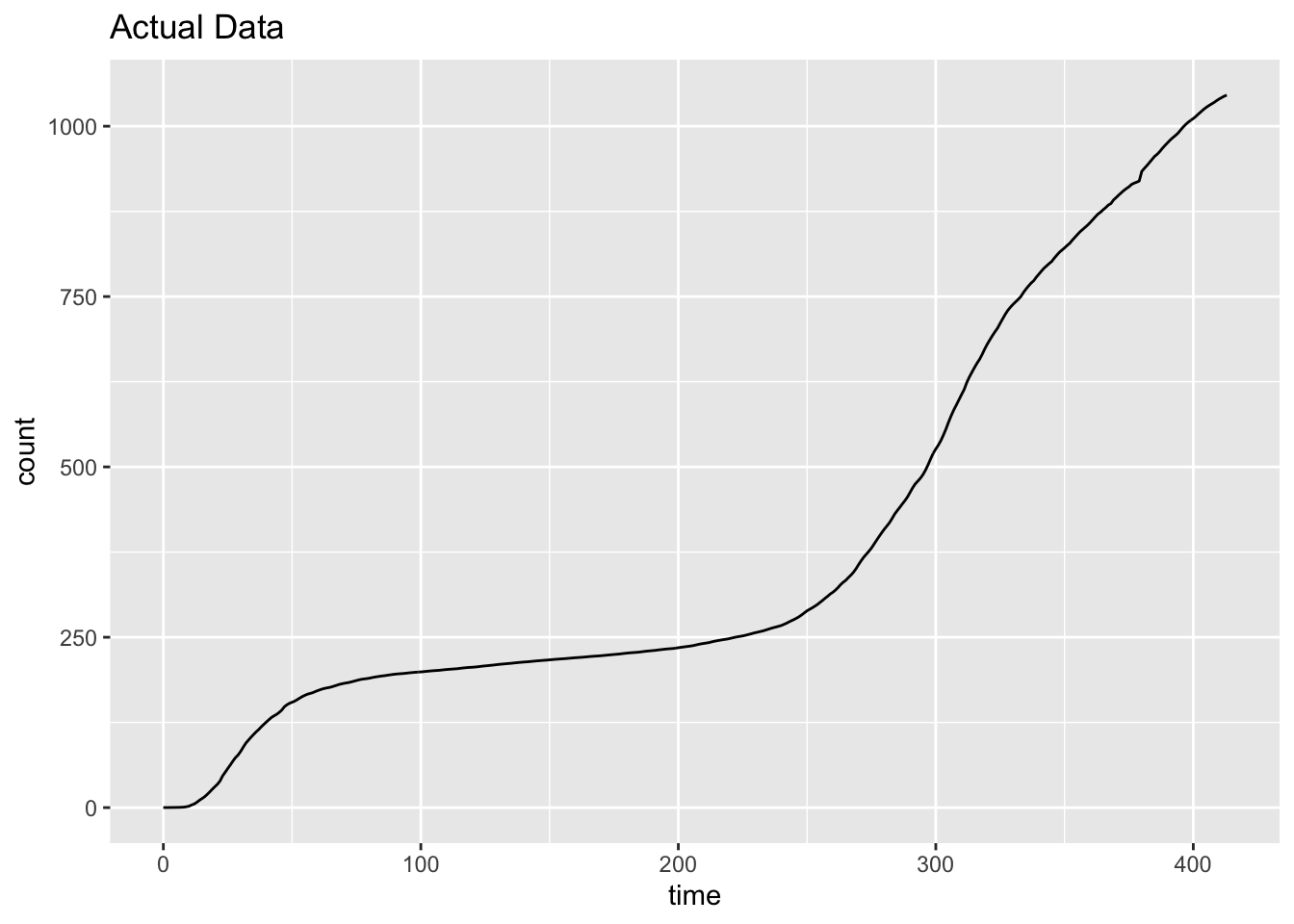
# redo for NY SSlogis
d <- 0:90
pred <- SSlogis(d, 186.233, 34.63, 9.4478)
test.1 <- data.frame(d = d, pred = c(NA, pred[-length(d)]),
pred.plus = c(NA, pred[-1]), group = rep(1, length(d)))
test.2 <- data.frame(d = d, pred = c(rep(NA, 2), pred[-seq(length(d) -
1, length(d), 1)]), pred.plus = c(rep(NA, 2), pred[-seq(1,
2, 1)]), group = rep(2, length(d)))
test.3 <- data.frame(d = d, pred = c(rep(NA, 3), pred[-seq(length(d) -
2, length(d), 1)]), pred.plus = c(rep(NA, 3), pred[-seq(1,
3, 1)]), group = rep(3, length(d)))
test.4 <- data.frame(d = d, pred = c(rep(NA, 4), pred[-seq(length(d) -
3, length(d), 1)]), pred.plus = c(rep(NA, 4), pred[-seq(1,
4, 1)]), group = rep(4, length(d)))
test.5 <- data.frame(d = d, pred = c(rep(NA, 5), pred[-seq(length(d) -
4, length(d), 1)]), pred.plus = c(rep(NA, 5), pred[-seq(1,
5, 1)]), group = rep(5, length(d)))
test <- rbind(test.1, test.2, test.3, test.4, test.5)
test$group <- factor(test$group)
ggplot(test, aes(x = pred, y = pred.plus, group = group,
color = group)) + geom_line() + xlab("f(t)") + ylab("f(t+k)") +
labs(color = "value of k")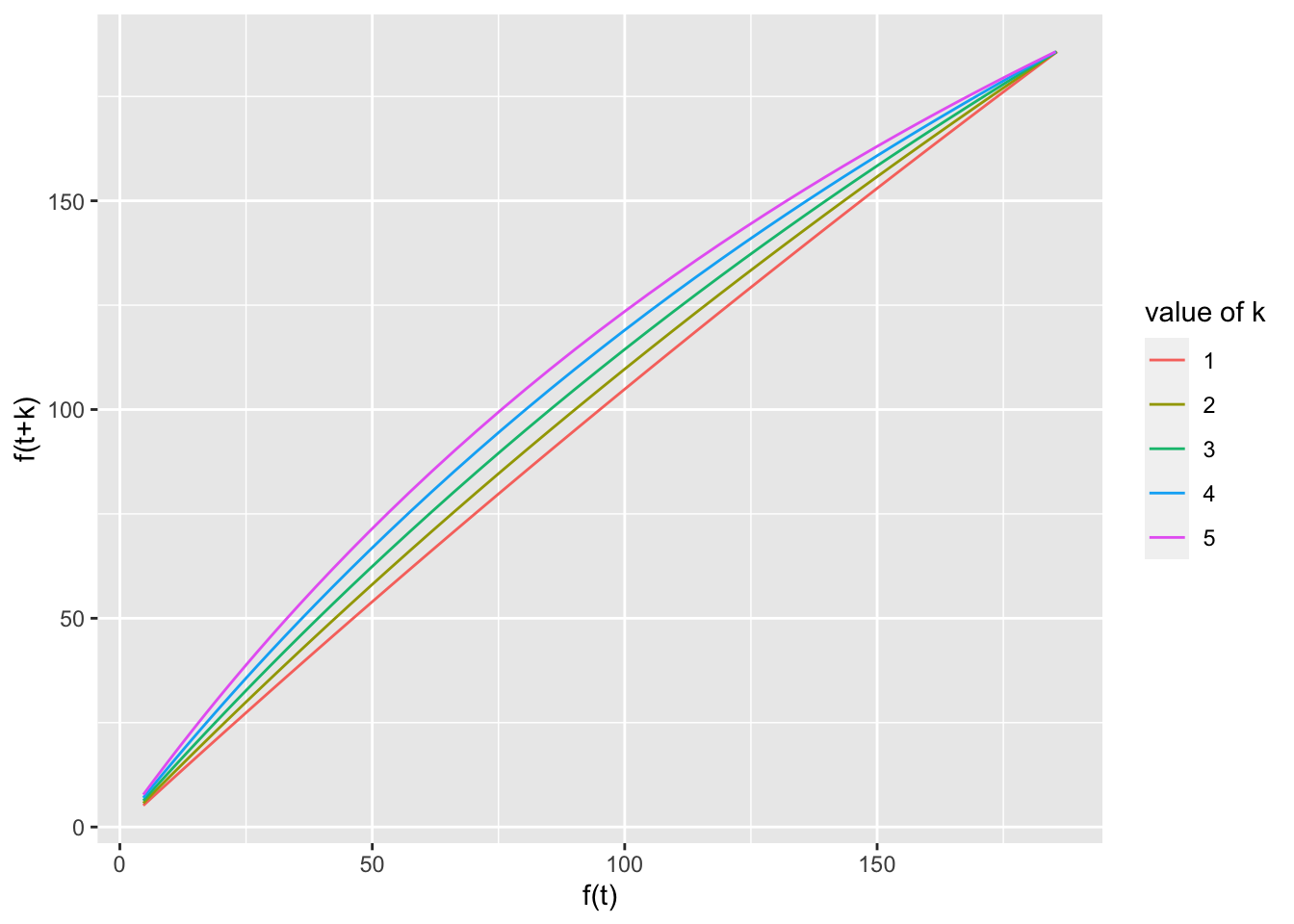
ggplot(data.frame(d, pred), aes(d, pred)) + geom_line() +
xlab("time") + ylab("count") + ggtitle("Best fitting logistic")